Similar analysis was found in Guangchuang Yu bioconductor vignettes page.
library(tidyverse);library(ShortRead);library(goseq);library(GO.db);library("annotate")## Warning: package 'S4Vectors' was built under R version 3.6.3## Warning: package 'GenomeInfoDb' was built under R version 3.6.3## Warning: package 'DelayedArray' was built under R version 3.6.3- Works only from campus network
At_cdna <- Biostrings::readDNAStringSet("https://www.arabidopsis.org/download_files/Sequences/TAIR10_blastsets/TAIR10_cds_20110103_representative_gene_model_updated") # In addition: Warning message:In download.file(fp, filepath2[i]) : URL 'https://www.arabidopsis.org/download_files/Sequences/TAIR10_blastsets/TAIR10_cds_20110103_representative_gene_model_updated': status was 'Peer certificate cannot be authenticated with given CA certificates' (032121). NO error from on campusUsing downloaded file (TAIR10_cdna_20110103_representative_gene_model_updated.gz)
getwd()## [1] "/Volumes/data_personal/Kazu_blog14/content/post/2018-09-27-over-representation-analysis-4-heatmap-visualization"system("gunzip -c ../../../resources/2018-09-26-over-representation-analysis-1-goseq-with-arabidopsis-go-term_files/TAIR10_cdna_20110103_representative_gene_model_updated.gz > ../../../resources/2018-09-26-over-representation-analysis-1-goseq-with-arabidopsis-go-term_files/TAIR10_cdna_20110103_representative_gene_model_updated")
At_cdna <- Biostrings::readDNAStringSet(file.path("..","..","..","resources","2018-09-26-over-representation-analysis-1-goseq-with-arabidopsis-go-term_files","TAIR10_cdna_20110103_representative_gene_model_updated"))
At_cdna## A DNAStringSet instance of length 33602
## width seq names
## [1] 2394 AGAAAACAGTCGACCGTCATT...TATTTGGTAATTTTTTGAGTC AT1G50920.1 | Sym...
## [2] 546 ATGACTCGTTTGTTGCCTTAC...CAAGTTGATTCTGGTACATAG AT1G36960.1 | Sym...
## [3] 2314 ATGGATTCAGAGTCAGAGTCA...CCTGGTGCATTGTGTTTCTCC AT1G44020.1 | Sym...
## [4] 1658 TCGTTTCGTCGTCGATCAGAA...ATCGATTACATGCTACATTTT AT1G15970.1 | Sym...
## [5] 1453 ATTGAAAAGAAAACACATCCC...GACCACCAAAATCTTCTCATA AT1G73440.1 | Sym...
## ... ... ...
## [33598] 87 GGATGGATGTCTGAGCGGTTG...GTTCGAATCCCTCTCCATCCG ATMG00420.1 | Sym...
## [33599] 384 ATGCTCCCCGCCGGTTGTTGG...CTACGATACTTAACTATATAA ATMG01330.1 | Sym...
## [33600] 573 ATGGATAACCAATTCATTTTC...GAACAGCGTAGCGACGGATAA ATMG00070.1 | Sym...
## [33601] 366 ATGGCATCAAAAATCCGCAAA...GATCCTTCTGCATACGCATAA ATMG00130.1 | Sym...
## [33602] 74 GCGCTCTTAGTTCAGTTCGGT...GTTCAAATCCTACAGAGCGTG ATMG00930.1 | Sym...# remove uncompressed file
system("rm ../../../resources/2018-09-26-over-representation-analysis-1-goseq-with-arabidopsis-go-term_files/TAIR10_cdna_20110103_representative_gene_model_updated")load GOseq function
# GOseq.Atgoslim.BP.list.ORA function
#This function is used to calculate GO ORA results multiple times.
getwd()## [1] "/Volumes/data_personal/Kazu_blog14/content/post/2018-09-27-over-representation-analysis-4-heatmap-visualization"# load("2018-09-27-over-representation-analysis-4-heatmap-visualization_files/Atgoslim.TAIR.BP.list.Rdata")
load("../../../resources/2018-09-26-over-representation-analysis-1-goseq-with-arabidopsis-go-term_files/Atgoslim.TAIR.BP.list.Rdata")
head(Atgoslim.TAIR.BP.list)## $`18S RRNA`
## [1] "GO:0006412"
##
## $`25S RRNA`
## [1] "GO:0043043" "GO:0043043" "GO:0043043"
##
## $`5.8S RRNA`
## [1] "GO:0043043" "GO:0043043" "GO:0043043"
##
## $AAN
## [1] "GO:0007623"
##
## $AAR1
## [1] "GO:0006970" "GO:0006970" "GO:0040029"
##
## $AAR2
## [1] "GO:0009845" "GO:0040029"GOseq.Atgoslim.BP.list.ORA<-function(genelist,padjust=0.05,ontology="BP",custom.category.list=Atgoslim.TAIR.BP.list,cdna=At_cdna) { # return GO enrichment table, padjus, padjust=0.05.
bias<-nchar(cdna)
names(bias) <- tibble(AGI=names(cdna)) %>% separate(AGI,into=c("AGI2" ,"test1","test2"),sep=" | ",extra="drop") %>% separate(AGI2,into="AGI3",sep="\\.",extra="drop") %>% dplyr::select(AGI3) %>% as_vector()
table(duplicated(names(bias))) # revised (032221)
#TF<-(names(bias) %in% genelist)*1
TF<-as.integer(names(bias) %in% genelist)
names(TF)<-names(bias)
#print(TF)
pwf<-nullp(TF,bias.data=bias)
#print(pwf$DEgenes)
GO.pval <- goseq(pwf,gene2cat=custom.category.list,use_genes_without_cat=TRUE) # format became different in new goseq version (021111). Does not work (042716)
#head(GO.pval)
if(ontology=="BP") {
GO.pval2<-subset(GO.pval,ontology=="BP")
} else if(ontology=="CC") {
GO.pval2<-subset(GO.pval,ontology=="CC")
} else {
GO.pval2<-subset(GO.pval,ontology=="MF")
}
# calculating padjust by BH
GO.pval2$over_represented_padjust<-p.adjust(GO.pval2$over_represented_pvalue,method="BH")
if(GO.pval2$over_represented_padjust[1]>padjust) return("no enriched GO")
else {
enriched.GO<-GO.pval2[GO.pval2$over_represented_padjust<padjust,]
print("enriched.GO is")
print(enriched.GO)
## write Term and Definition
for(i in 1:dim(enriched.GO)[1]) {
if(is.null(Term(GOTERM[enriched.GO[i,"category"]]))) {next} else {
enriched.GO$Term[i]<-Term(GOTERM[[enriched.GO[i,"category"]]])
enriched.GO$Definition[i]<-Definition(GOTERM[[enriched.GO[i,"category"]]])
}
}
return(enriched.GO)
}
}Importing enrichment result table and have summary table: Table of DEGs (under construction)
# Ding (2018) Cell DEGs
getwd()## [1] "/Volumes/data_personal/Kazu_blog14/content/post/2018-09-27-over-representation-analysis-4-heatmap-visualization"DEG.objs <- list.files(path=file.path("..","..","..","resources","2018-09-27-over-representation-analysis-4-heatmap-visualization_files","output"),pattern="\\.csv$",full.names=TRUE) #
DEG.objs2 <- list.files(path=file.path("..","..","..","resources","2018-09-27-over-representation-analysis-4-heatmap-visualization_files","output"),pattern="\\.csv$") #
# effect of "full.names=TRUE/FALSE"
DEG.objs## [1] "../../../resources/2018-09-27-over-representation-analysis-4-heatmap-visualization_files/output/genotypenpr1_genotypenpr1.treatmenttreated.DEGs.int.rWT.rU.csv"
## [2] "../../../resources/2018-09-27-over-representation-analysis-4-heatmap-visualization_files/output/genotypenpr4_genotypenpr4.treatmenttreated.DEGs.int.rWT.rU.csv"
## [3] "../../../resources/2018-09-27-over-representation-analysis-4-heatmap-visualization_files/output/trt.DEGs.int.rWT.rU.csv"# read csv file
DEG.map <- DEG.objs %>% map(.,function(x) read_csv(x))
# or
DEG.map <- DEG.objs %>% map(~read_csv(.))
# name each data.frame
names(DEG.map) <- as_vector(gsub(".csv","",DEG.objs2))
DEG.map## $genotypenpr1_genotypenpr1.treatmenttreated.DEGs.int.rWT.rU
## # A tibble: 6,694 x 10
## genes logFC.genotypen… logFC.genotypen… logCPM LR PValue FDR
## <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 AT2G… -4.20 -0.720 6.69 1300. 5.30e-283 1.78e-278
## 2 AT5G… 3.26 0.530 4.76 1000. 5.96e-218 1.00e-213
## 3 AT1G… -2.02 -0.475 7.19 629. 2.69e-137 3.01e-133
## 4 AT1G… -2.26 -0.511 7.51 620. 2.83e-135 2.38e-131
## 5 AT3G… -3.16 0.888 5.33 618. 5.42e-135 3.64e-131
## 6 AT5G… -0.212 -6.72 3.89 607. 1.67e-132 9.35e-129
## 7 AT2G… 2.22 1.95 9.21 572. 7.80e-125 3.74e-121
## 8 AT5G… 1.82 1.83 9.40 570. 1.54e-124 6.46e-121
## 9 AT1G… -3.65 -1.52 3.37 565. 1.92e-123 7.17e-120
## 10 AT5G… 0.000149 -7.96 3.77 546. 2.42e-119 8.12e-116
## # … with 6,684 more rows, and 3 more variables: AGI <chr>, symbol2 <chr>,
## # full_name <chr>
##
## $genotypenpr4_genotypenpr4.treatmenttreated.DEGs.int.rWT.rU
## # A tibble: 846 x 10
## genes logFC.genotypen… logFC.genotypen… logCPM LR PValue FDR
## <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 AT5G… 3.04 0.628 4.76 904. 5.00e-197 1.68e-192
## 2 AT3G… -5.45 -0.183 1.98 459. 1.80e-100 3.03e- 96
## 3 AT5G… -3.30 0.354 3.10 444. 4.51e- 97 5.05e- 93
## 4 AT1G… -7.40 -3.70 2.00 432. 1.32e- 94 1.11e- 90
## 5 AT3G… -2.97 -0.301 3.36 372. 1.83e- 81 1.23e- 77
## 6 AT1G… -2.50 0.891 6.30 288. 2.54e- 63 1.42e- 59
## 7 AT2G… -1.42 -0.703 6.69 277. 7.63e- 61 3.66e- 57
## 8 AT5G… -3.04 0.434 4.19 211. 1.75e- 46 7.36e- 43
## 9 AT2G… -1.10 -0.237 5.87 209. 5.09e- 46 1.90e- 42
## 10 AT5G… -1.84 -0.489 2.78 187. 2.68e- 41 9.01e- 38
## # … with 836 more rows, and 3 more variables: AGI <chr>, symbol2 <chr>,
## # full_name <chr>
##
## $trt.DEGs.int.rWT.rU
## # A tibble: 7,811 x 9
## genes logFC logCPM LR PValue FDR AGI symbol2 full_name
## <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <chr> <chr> <chr>
## 1 AT5G1… 8.57 3.89 675. 7.55e-149 2.54e-144 AT5G1… AT-HSP1… heat shock pro…
## 2 AT1G0… 4.00 4.60 592. 7.94e-131 1.33e-126 AT1G0… <NA> <NA>
## 3 AT5G2… 6.16 4.19 588. 5.77e-130 6.46e-126 AT5G2… ATWRKY3… ARABIDOPSIS TH…
## 4 AT4G2… 3.85 5.39 586. 1.92e-129 1.59e-125 AT4G2… CRK14 cysteine-rich …
## 5 AT5G1… 9.64 3.77 586. 2.37e-129 1.59e-125 AT5G1… HSP17.6… 17.6 kDa class…
## 6 AT3G2… 4.92 3.36 546. 9.68e-121 5.42e-117 AT3G2… <NA> <NA>
## 7 AT4G2… 3.34 6.70 539. 2.78e-119 1.33e-115 AT4G2… SGT1A <NA>
## 8 AT5G2… 3.82 6.70 535. 2.42e-118 1.02e-114 AT5G2… AtDMR6;… NA;DOWNY MILDE…
## 9 AT5G0… 4.14 7.14 523. 1.07e-115 3.98e-112 AT5G0… AtHsp70… NA;NA
## 10 AT1G2… 2.87 7.19 511. 3.43e-113 1.15e-109 AT1G2… AtWAK1;… NA;NA;cell wal…
## # … with 7,801 more rows# GOseq.Atgoslim.BP.list.ORA(genelist=genes.shade1h.up$AGI)GO.ORA for each DEG list (loop)
# single coefficient
names(DEG.map)## [1] "genotypenpr1_genotypenpr1.treatmenttreated.DEGs.int.rWT.rU"
## [2] "genotypenpr4_genotypenpr4.treatmenttreated.DEGs.int.rWT.rU"
## [3] "trt.DEGs.int.rWT.rU"for(n in 3:3) {
library(dplyr)
#genelist.up<-base::get(paste("../output/",DEG.objs.V1.5annotation.unique[n],sep="")) %>% rownames_to_column() %>% dplyr::filter(logFC>0) # does not work
genelist.up <- DEG.map[[n]] %>% dplyr::filter(logFC>0)
genelist.down <-DEG.map[[n]] %>% dplyr::filter(logFC<0)
GO.ORA.temp.up<-GOseq.Atgoslim.BP.list.ORA(str_remove(as_vector(genelist.up[,"genes"]),"\\.[[:digit:]]+"),custom.category.list=Atgoslim.TAIR.BP.list)
GO.ORA.temp.down<-GOseq.Atgoslim.BP.list.ORA(str_remove(as_vector(genelist.down[,"genes"]),"\\.[[:digit:]]+"),custom.category.list=Atgoslim.TAIR.BP.list)
# handling "no enriched GO"
# genelist.names<-c("GO.ORA.temp.up_down","GO.ORA.temp.down_up") # test
x<-list(GO.ORA.temp.up=GO.ORA.temp.up,
GO.ORA.temp.down=GO.ORA.temp.down) # list
x<-x[!x=="no enriched GO"] # reove "no enriched GO" result
## add sample info and FC info and save GO.ORA result
for (i in 1:length(x)) {
GO.ORA.result<-x[[i]] %>% mutate(FC = gsub("(GO.ORA.temp.)(.)","\\2",names(x)[i]),sample=DEG.objs[n])
save(GO.ORA.result,file=file.path("output",paste(gsub(".csv","",DEG.objs2[n]),gsub("(GO.ORA.temp.)(.)","\\2",names(x)[i]),"enrich.Rdata",sep=".")))
rm(GO.ORA.result)
}
}## Warning in pcls(G): initial point very close to some inequality constraints## [1] "enriched.GO is"
## category over_represented_pvalue under_represented_pvalue numDEInCat
## 3214 GO:0071456 3.522448e-38 1.0000000 111
## 2314 GO:0042742 3.716595e-32 1.0000000 151
## 638 GO:0006886 8.866593e-32 1.0000000 98
## 444 GO:0006468 9.412221e-31 1.0000000 281
## 737 GO:0007165 1.305373e-23 1.0000000 138
## 1227 GO:0010200 4.324664e-22 1.0000000 65
## 919 GO:0009617 8.090698e-20 1.0000000 58
## 997 GO:0009751 1.120259e-19 1.0000000 54
## 640 GO:0006888 4.058569e-19 1.0000000 45
## 875 GO:0009408 1.464687e-17 1.0000000 68
## 2824 GO:0050832 2.346560e-17 1.0000000 80
## 1603 GO:0016192 7.596859e-16 1.0000000 59
## 915 GO:0009611 1.640827e-15 1.0000000 75
## 945 GO:0009651 9.974609e-14 1.0000000 111
## 2654 GO:0046777 1.559745e-13 1.0000000 71
## 3367 GO:0080167 1.182305e-12 1.0000000 49
## 920 GO:0009620 9.339184e-12 1.0000000 41
## 642 GO:0006891 1.351765e-11 1.0000000 24
## 985 GO:0009737 3.967797e-11 1.0000000 110
## 641 GO:0006890 6.040677e-11 1.0000000 21
## 880 GO:0009414 8.718892e-11 1.0000000 86
## 201 GO:0002239 2.782784e-10 1.0000000 17
## 199 GO:0002237 5.337928e-10 1.0000000 21
## 441 GO:0006457 4.312064e-09 1.0000000 45
## 923 GO:0009626 6.127232e-09 1.0000000 24
## 1611 GO:0016310 9.371211e-09 1.0000000 77
## 690 GO:0007030 9.885730e-09 1.0000000 20
## 658 GO:0006952 3.410617e-08 1.0000000 52
## 2869 GO:0051259 3.491474e-08 1.0000000 11
## 1863 GO:0030968 8.668916e-08 1.0000000 17
## 876 GO:0009409 9.034140e-08 1.0000000 83
## 2938 GO:0051707 1.159197e-07 1.0000000 15
## 2141 GO:0034605 1.679920e-07 1.0000000 24
## 738 GO:0007166 1.817005e-07 1.0000000 22
## 3586 GO:1900150 2.006935e-07 1.0000000 18
## 2757 GO:0048544 2.010186e-07 1.0000000 20
## 200 GO:0002238 2.755361e-07 1.0000000 12
## 2207 GO:0035556 2.816263e-07 0.9999999 56
## 628 GO:0006874 6.212941e-07 0.9999999 19
## 2512 GO:0045332 6.780943e-07 1.0000000 10
## 1894 GO:0031204 7.203511e-07 1.0000000 8
## 865 GO:0009306 8.660987e-07 0.9999999 12
## 2299 GO:0042542 1.065578e-06 0.9999998 23
## 3507 GO:0098542 1.526214e-06 0.9999997 19
## 1051 GO:0009862 2.049848e-06 0.9999998 12
## 2497 GO:0045087 2.590178e-06 0.9999993 25
## 3133 GO:0070973 3.587809e-06 0.9999998 8
## 3881 GO:2000022 3.853147e-06 0.9999992 18
## 462 GO:0006499 4.401279e-06 0.9999991 19
## 874 GO:0009407 4.497272e-06 0.9999991 17
## 3280 GO:0072583 5.737080e-06 0.9999989 17
## 924 GO:0009627 5.994178e-06 0.9999986 20
## 1052 GO:0009863 6.405782e-06 0.9999991 13
## 197 GO:0002229 6.854004e-06 0.9999980 27
## 3601 GO:1900426 6.870626e-06 0.9999990 14
## 665 GO:0006979 7.517450e-06 0.9999964 67
## 1201 GO:0010150 8.786798e-06 0.9999968 34
## 2646 GO:0046686 9.690411e-06 0.9999951 74
## 1222 GO:0010193 1.025804e-05 0.9999982 15
## 647 GO:0006897 1.665257e-05 0.9999961 19
## 443 GO:0006465 1.801892e-05 0.9999984 9
## 3348 GO:0080142 1.987307e-05 0.9999986 9
## 605 GO:0006816 2.086469e-05 0.9999959 16
## 1246 GO:0010224 2.306386e-05 0.9999933 23
## 1667 GO:0018105 2.378760e-05 0.9999914 32
## 998 GO:0009753 2.520894e-05 0.9999905 33
## 2262 GO:0042147 2.675004e-05 0.9999952 14
## 661 GO:0006970 3.172258e-05 0.9999865 40
## 2281 GO:0042335 3.926545e-05 0.9999933 13
## 196 GO:0002221 3.981563e-05 0.9999965 9
## 2144 GO:0034620 4.446797e-05 0.9999951 10
## 1185 GO:0010112 5.121833e-05 0.9999959 8
## 3274 GO:0072334 6.035692e-05 0.9999986 6
## 917 GO:0009615 8.111631e-05 0.9999809 16
## 575 GO:0006749 8.456719e-05 0.9999747 20
## 2520 GO:0045489 8.474236e-05 0.9999815 15
## 2302 GO:0042546 1.800576e-04 0.9999475 18
## 3041 GO:0061025 1.993975e-04 0.9999606 12
## 1899 GO:0031347 1.999445e-04 0.9999419 18
## 2707 GO:0048278 2.092712e-04 0.9999621 11
## 3355 GO:0080151 2.241618e-04 0.9999944 5
## 1844 GO:0030433 2.263682e-04 0.9999406 16
## 2969 GO:0052544 2.374330e-04 0.9999699 9
## 3600 GO:1900425 2.392471e-04 0.9999683 9
## 3030 GO:0060866 2.418957e-04 0.9999882 6
## 1905 GO:0031408 2.817122e-04 0.9999618 9
## 2385 GO:0043207 3.106536e-04 1.0000000 3
## 206 GO:0002758 3.198846e-04 0.9999409 11
## 771 GO:0008219 3.208565e-04 0.9999284 13
## 1056 GO:0009867 3.514213e-04 0.9998875 19
## 2823 GO:0050829 3.759602e-04 0.9999366 10
## 1132 GO:0010038 4.144226e-04 0.9999463 8
## 990 GO:0009742 4.471442e-04 0.9998483 20
## 741 GO:0007186 4.564777e-04 0.9999423 8
## 652 GO:0006906 4.576189e-04 0.9999062 11
## 3191 GO:0071323 4.593066e-04 0.9999864 5
## 2952 GO:0051865 4.852158e-04 0.9999261 9
## 3398 GO:0090114 4.994257e-04 1.0000000 4
## 1193 GO:0010120 5.345849e-04 0.9999453 7
## 639 GO:0006887 5.711489e-04 0.9998339 16
## 1454 GO:0010942 5.875836e-04 0.9999577 6
## 2123 GO:0034440 6.183469e-04 0.9999804 5
## 2840 GO:0051085 6.784697e-04 0.9997954 16
## 1318 GO:0010359 6.904112e-04 1.0000000 4
## 694 GO:0007034 6.982514e-04 0.9998817 9
## 1288 GO:0010286 7.474543e-04 0.9997697 16
## 3107 GO:0070588 8.083288e-04 0.9997953 13
## 646 GO:0006896 8.367341e-04 0.9998168 11
## 1900 GO:0031348 8.477306e-04 0.9998165 11
## 648 GO:0006898 1.046236e-03 0.9999151 6
## 3404 GO:0090158 1.091496e-03 0.9998903 6
## 2678 GO:0047484 1.137873e-03 0.9998508 7
## 651 GO:0006904 1.167987e-03 0.9998533 7
## 3397 GO:0090110 1.172567e-03 0.9998616 7
## 3169 GO:0071230 1.344001e-03 0.9997235 10
## numInCat
## 3214 238
## 2314 391
## 638 204
## 444 928
## 737 411
## 1227 138
## 919 118
## 997 108
## 640 85
## 875 169
## 2824 218
## 1603 153
## 915 209
## 945 395
## 2654 188
## 3367 128
## 920 99
## 642 40
## 985 423
## 641 34
## 880 318
## 201 24
## 199 34
## 441 138
## 923 46
## 1611 273
## 690 37
## 658 192
## 2869 17
## 1863 31
## 876 349
## 2938 24
## 2141 59
## 738 46
## 3586 34
## 2757 38
## 200 17
## 2207 185
## 628 37
## 2512 12
## 1894 10
## 865 19
## 2299 62
## 3507 44
## 1051 19
## 2497 68
## 3133 11
## 3881 42
## 462 46
## 874 46
## 3280 36
## 924 54
## 1052 25
## 197 74
## 3601 27
## 665 301
## 1201 115
## 2646 312
## 1222 34
## 647 45
## 443 16
## 3348 13
## 605 34
## 1246 67
## 1667 98
## 998 113
## 2262 31
## 661 152
## 2281 28
## 196 14
## 2144 17
## 1185 13
## 3274 7
## 917 40
## 575 68
## 2520 35
## 2302 56
## 3041 29
## 1899 53
## 2707 26
## 3355 6
## 1844 42
## 2969 16
## 3600 17
## 3030 8
## 1905 17
## 2385 3
## 206 24
## 771 31
## 1056 59
## 2823 21
## 1132 16
## 990 64
## 741 15
## 652 28
## 3191 6
## 2952 18
## 3398 4
## 1193 12
## 639 44
## 1454 9
## 2123 6
## 2840 48
## 1318 4
## 694 20
## 1288 51
## 3107 33
## 646 27
## 1900 26
## 648 9
## 3404 11
## 2678 14
## 651 13
## 3397 12
## 3169 22
## term
## 3214 cellular response to hypoxia
## 2314 defense response to bacterium
## 638 intracellular protein transport
## 444 protein phosphorylation
## 737 signal transduction
## 1227 response to chitin
## 919 response to bacterium
## 997 response to salicylic acid
## 640 ER to Golgi vesicle-mediated transport
## 875 response to heat
## 2824 defense response to fungus
## 1603 vesicle-mediated transport
## 915 response to wounding
## 945 response to salt stress
## 2654 protein autophosphorylation
## 3367 response to karrikin
## 920 response to fungus
## 642 intra-Golgi vesicle-mediated transport
## 985 response to abscisic acid
## 641 retrograde vesicle-mediated transport, Golgi to ER
## 880 response to water deprivation
## 201 response to oomycetes
## 199 response to molecule of bacterial origin
## 441 protein folding
## 923 plant-type hypersensitive response
## 1611 phosphorylation
## 690 Golgi organization
## 658 defense response
## 2869 protein complex oligomerization
## 1863 endoplasmic reticulum unfolded protein response
## 876 response to cold
## 2938 response to other organism
## 2141 cellular response to heat
## 738 cell surface receptor signaling pathway
## 3586 regulation of defense response to fungus
## 2757 recognition of pollen
## 200 response to molecule of fungal origin
## 2207 intracellular signal transduction
## 628 cellular calcium ion homeostasis
## 2512 phospholipid translocation
## 1894 posttranslational protein targeting to membrane, translocation
## 865 protein secretion
## 2299 response to hydrogen peroxide
## 3507 defense response to other organism
## 1051 systemic acquired resistance, salicylic acid mediated signaling pathway
## 2497 innate immune response
## 3133 protein localization to endoplasmic reticulum exit site
## 3881 regulation of jasmonic acid mediated signaling pathway
## 462 N-terminal protein myristoylation
## 874 toxin catabolic process
## 3280 clathrin-dependent endocytosis
## 924 systemic acquired resistance
## 1052 salicylic acid mediated signaling pathway
## 197 defense response to oomycetes
## 3601 positive regulation of defense response to bacterium
## 665 response to oxidative stress
## 1201 leaf senescence
## 2646 response to cadmium ion
## 1222 response to ozone
## 647 endocytosis
## 443 signal peptide processing
## 3348 regulation of salicylic acid biosynthetic process
## 605 calcium ion transport
## 1246 response to UV-B
## 1667 peptidyl-serine phosphorylation
## 998 response to jasmonic acid
## 2262 retrograde transport, endosome to Golgi
## 661 response to osmotic stress
## 2281 cuticle development
## 196 pattern recognition receptor signaling pathway
## 2144 cellular response to unfolded protein
## 1185 regulation of systemic acquired resistance
## 3274 UDP-galactose transmembrane transport
## 917 response to virus
## 575 glutathione metabolic process
## 2520 pectin biosynthetic process
## 2302 cell wall biogenesis
## 3041 membrane fusion
## 1899 regulation of defense response
## 2707 vesicle docking
## 3355 positive regulation of salicylic acid mediated signaling pathway
## 1844 ubiquitin-dependent ERAD pathway
## 2969 defense response by callose deposition in cell wall
## 3600 negative regulation of defense response to bacterium
## 3030 leaf abscission
## 1905 oxylipin biosynthetic process
## 2385 response to external biotic stimulus
## 206 innate immune response-activating signal transduction
## 771 cell death
## 1056 jasmonic acid mediated signaling pathway
## 2823 defense response to Gram-negative bacterium
## 1132 response to metal ion
## 990 brassinosteroid mediated signaling pathway
## 741 G protein-coupled receptor signaling pathway
## 652 vesicle fusion
## 3191 cellular response to chitin
## 2952 protein autoubiquitination
## 3398 COPII-coated vesicle budding
## 1193 camalexin biosynthetic process
## 639 exocytosis
## 1454 positive regulation of cell death
## 2123 lipid oxidation
## 2840 chaperone cofactor-dependent protein refolding
## 1318 regulation of anion channel activity
## 694 vacuolar transport
## 1288 heat acclimation
## 3107 calcium ion transmembrane transport
## 646 Golgi to vacuole transport
## 1900 negative regulation of defense response
## 648 receptor-mediated endocytosis
## 3404 endoplasmic reticulum membrane organization
## 2678 regulation of response to osmotic stress
## 651 vesicle docking involved in exocytosis
## 3397 cargo loading into COPII-coated vesicle
## 3169 cellular response to amino acid stimulus
## ontology over_represented_padjust
## 3214 BP 1.394537e-34
## 2314 BP 7.357000e-29
## 638 BP 1.170095e-28
## 444 BP 9.315746e-28
## 737 BP 1.033594e-20
## 1227 BP 2.853557e-19
## 919 BP 4.575868e-17
## 997 BP 5.543881e-17
## 640 BP 1.785320e-16
## 875 BP 5.798697e-15
## 2824 BP 8.445485e-15
## 1603 BP 2.506330e-13
## 915 BP 4.996951e-13
## 945 BP 2.820677e-11
## 2654 BP 4.116688e-11
## 3367 BP 2.925467e-10
## 920 BP 2.174931e-09
## 642 BP 2.973132e-09
## 985 BP 8.267636e-09
## 641 BP 1.195752e-08
## 880 BP 1.643719e-08
## 201 BP 5.007746e-08
## 199 BP 9.188198e-08
## 441 BP 7.113109e-07
## 923 BP 9.703084e-07
## 1611 BP 1.426947e-06
## 690 BP 1.449541e-06
## 658 BP 4.766464e-06
## 2869 BP 4.766464e-06
## 1863 BP 1.144008e-05
## 876 BP 1.153747e-05
## 2938 BP 1.434144e-05
## 2141 BP 2.015395e-05
## 738 BP 2.115742e-05
## 3586 BP 2.210646e-05
## 2757 BP 2.210646e-05
## 200 BP 2.934102e-05
## 2207 BP 2.934102e-05
## 628 BP 6.306932e-05
## 2512 BP 6.711438e-05
## 1894 BP 6.955781e-05
## 865 BP 8.164011e-05
## 2299 BP 9.810750e-05
## 3507 BP 1.373246e-04
## 1051 BP 1.803411e-04
## 2497 BP 2.229243e-04
## 3133 BP 3.022156e-04
## 3881 BP 3.178044e-04
## 462 BP 3.556054e-04
## 874 BP 3.560940e-04
## 3280 BP 4.453549e-04
## 924 BP 4.563645e-04
## 1052 BP 4.784998e-04
## 197 BP 4.945601e-04
## 3601 BP 4.945601e-04
## 665 BP 5.314569e-04
## 1201 BP 6.102971e-04
## 2646 BP 6.614541e-04
## 1222 BP 6.883317e-04
## 647 BP 1.098792e-03
## 443 BP 1.169458e-03
## 3348 BP 1.268992e-03
## 605 BP 1.311164e-03
## 1246 BP 1.426716e-03
## 1667 BP 1.448848e-03
## 998 BP 1.512154e-03
## 2262 BP 1.580648e-03
## 661 BP 1.846907e-03
## 2281 BP 2.251858e-03
## 196 BP 2.251858e-03
## 2144 BP 2.479559e-03
## 1185 BP 2.816297e-03
## 3274 BP 3.273330e-03
## 917 BP 4.339723e-03
## 575 BP 4.414408e-03
## 2520 BP 4.414408e-03
## 2302 BP 9.257769e-03
## 3041 BP 1.002000e-02
## 1899 BP 1.002000e-02
## 2707 BP 1.035631e-02
## 3355 BP 1.092917e-02
## 1844 BP 1.092917e-02
## 2969 BP 1.126665e-02
## 3600 BP 1.126665e-02
## 3030 BP 1.126665e-02
## 1905 BP 1.296859e-02
## 2385 BP 1.413653e-02
## 206 BP 1.427271e-02
## 771 BP 1.427271e-02
## 1056 BP 1.545863e-02
## 2823 BP 1.635634e-02
## 1132 BP 1.783368e-02
## 990 BP 1.894161e-02
## 741 BP 1.894161e-02
## 652 BP 1.894161e-02
## 3191 BP 1.894161e-02
## 2952 BP 1.980381e-02
## 3398 BP 2.017578e-02
## 1193 BP 2.137800e-02
## 639 BP 2.261179e-02
## 1454 BP 2.303212e-02
## 2123 BP 2.400035e-02
## 2840 BP 2.607827e-02
## 1318 BP 2.628210e-02
## 694 BP 2.632740e-02
## 1288 BP 2.791671e-02
## 3107 BP 2.990817e-02
## 646 BP 3.067250e-02
## 1900 BP 3.079051e-02
## 648 BP 3.765499e-02
## 3404 BP 3.893002e-02
## 2678 BP 4.022177e-02
## 651 BP 4.072100e-02
## 3397 BP 4.072100e-02
## 3169 BP 4.626869e-02## Warning in pcls(G): initial point very close to some inequality constraints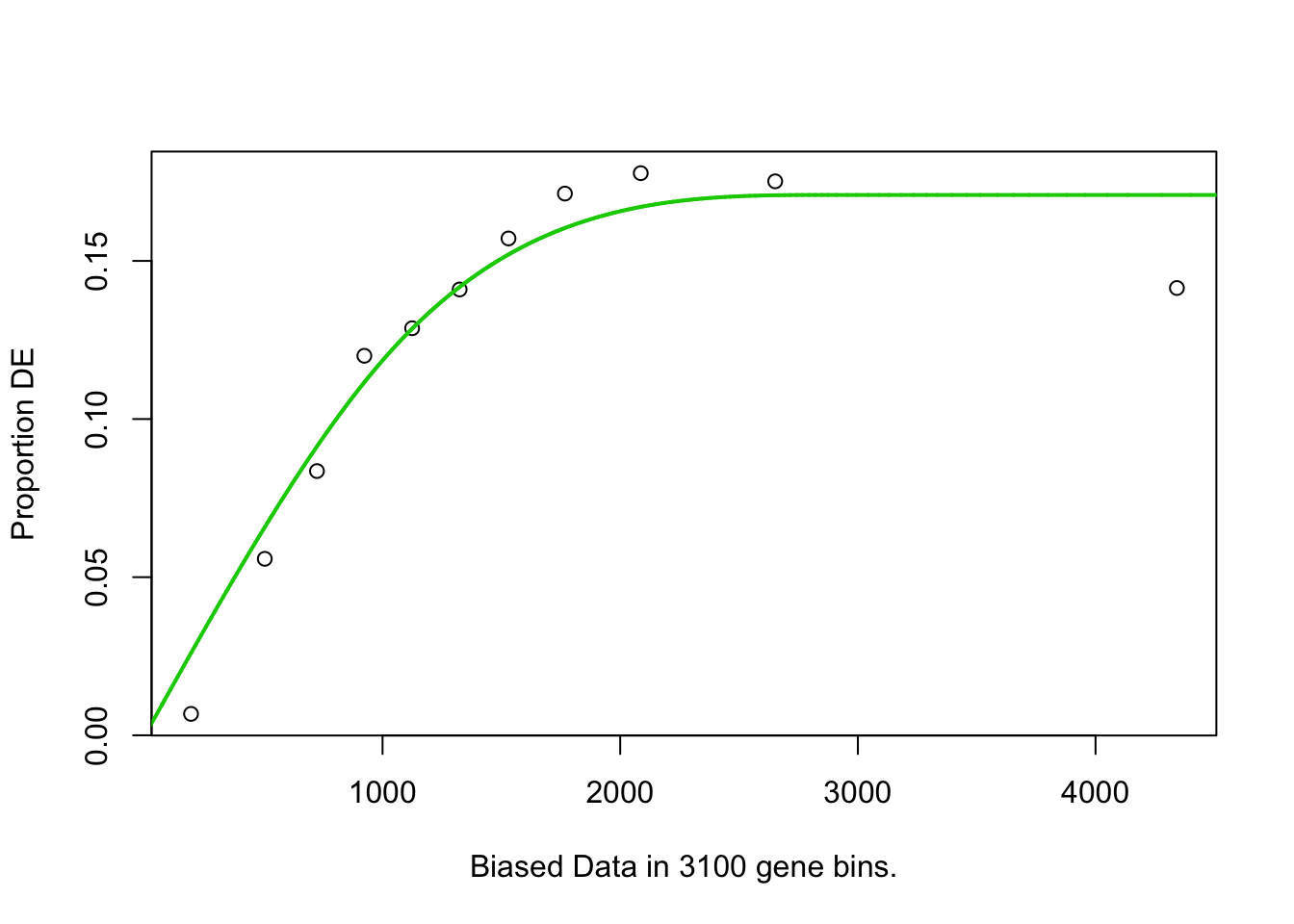
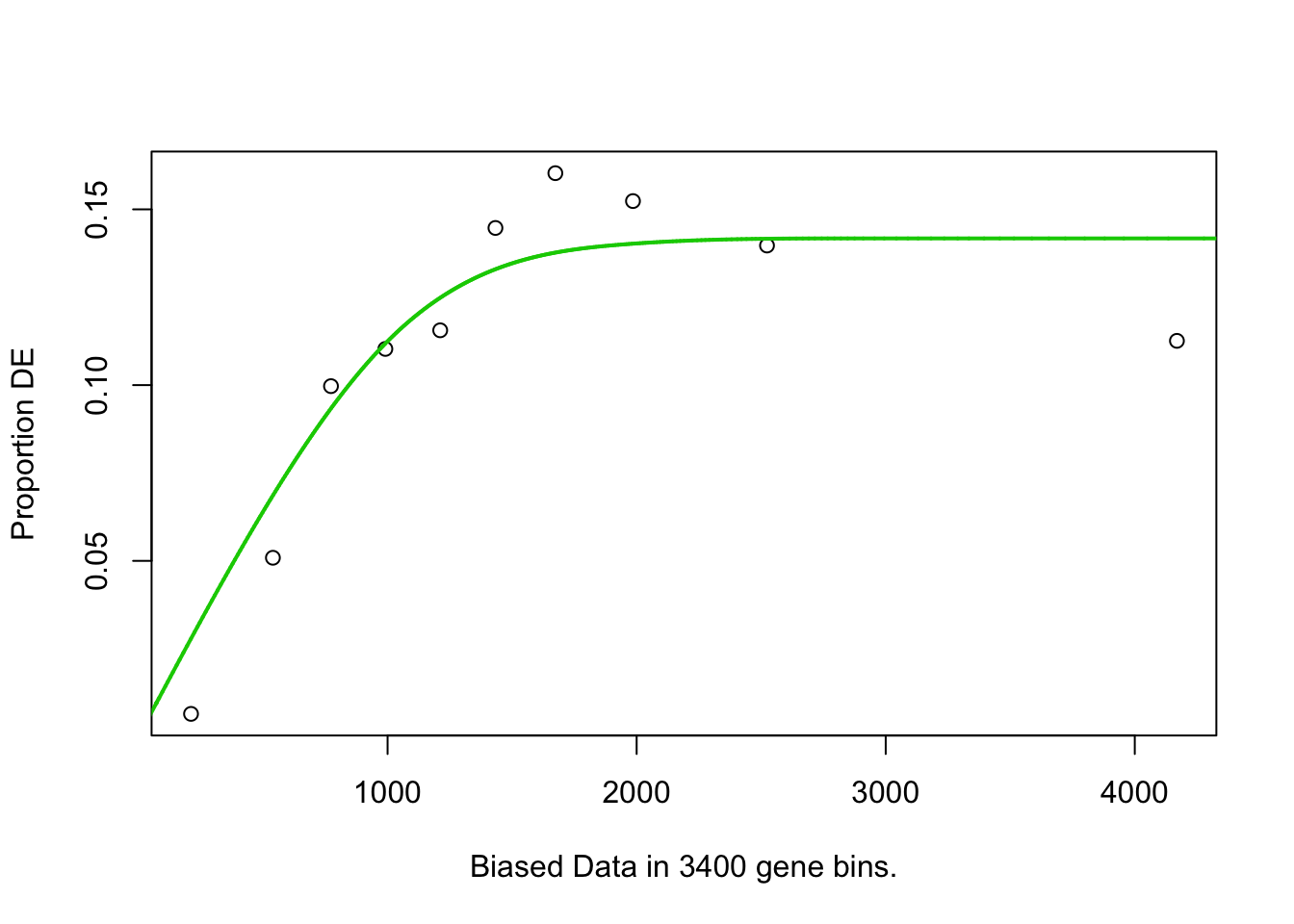
## [1] "enriched.GO is"
## category over_represented_pvalue under_represented_pvalue numDEInCat
## 949 GO:0009658 1.357052e-30 1.0000000 87
## 882 GO:0009416 6.448020e-15 1.0000000 69
## 1003 GO:0009768 2.442671e-14 1.0000000 19
## 3625 GO:1901259 3.405670e-14 1.0000000 18
## 1556 GO:0015979 2.009362e-12 1.0000000 29
## 1560 GO:0015995 1.330935e-11 1.0000000 24
## 1123 GO:0010027 1.439752e-10 1.0000000 25
## 1273 GO:0010258 3.818474e-09 1.0000000 8
## 2333 GO:0042793 1.206797e-08 1.0000000 10
## 760 GO:0007623 1.229726e-08 1.0000000 36
## 1006 GO:0009773 2.085945e-08 1.0000000 12
## 939 GO:0009645 3.512489e-08 1.0000000 9
## 379 GO:0006355 3.598346e-08 1.0000000 238
## 2486 GO:0045036 8.592642e-08 1.0000000 13
## 876 GO:0009409 1.122844e-07 1.0000000 76
## 1258 GO:0010239 1.260398e-07 1.0000000 9
## 938 GO:0009644 2.509688e-07 1.0000000 19
## 1223 GO:0010196 7.163210e-07 1.0000000 10
## 1233 GO:0010207 9.303939e-07 0.9999999 13
## 940 GO:0009646 2.151375e-06 0.9999996 17
## 2487 GO:0045037 2.666368e-06 0.9999996 14
## 683 GO:0007018 4.477986e-06 0.9999988 23
## 948 GO:0009657 4.623809e-06 0.9999997 9
## 1187 GO:0010114 4.820165e-06 0.9999989 20
## 1116 GO:0010020 5.985844e-06 0.9999993 12
## 3225 GO:0071482 7.932741e-06 0.9999996 8
## 1002 GO:0009767 1.131163e-05 0.9999988 10
## 227 GO:0005983 1.273424e-05 0.9999988 10
## 1906 GO:0031425 1.525556e-05 0.9999988 9
## 931 GO:0009637 2.556168e-05 0.9999938 18
## 1590 GO:0016120 3.279494e-05 0.9999993 6
## 1117 GO:0010021 4.002085e-05 0.9999991 6
## 1015 GO:0009793 4.384335e-05 0.9999745 89
## 69 GO:0000373 4.730942e-05 0.9999924 12
## 2722 GO:0048366 5.612080e-05 0.9999771 34
## 1004 GO:0009769 7.064768e-05 0.9999962 6
## 932 GO:0009638 9.873746e-05 0.9999842 11
## 146 GO:0000967 1.049452e-04 0.9999978 5
## 1001 GO:0009765 1.096343e-04 0.9999977 5
## 1593 GO:0016123 1.102692e-04 0.9999954 6
## 3607 GO:1900865 1.580975e-04 0.9999757 10
## 3608 GO:1900871 1.589997e-04 1.0000000 4
## 1167 GO:0010087 1.755620e-04 0.9999483 18
## 1703 GO:0019252 1.760930e-04 0.9999592 14
## 987 GO:0009739 2.077467e-04 0.9999325 20
## 375 GO:0006351 2.386003e-04 0.9999096 25
## 238 GO:0006002 2.708948e-04 0.9999685 8
## 1993 GO:0032544 2.919623e-04 0.9999548 9
## 2411 GO:0043489 2.959193e-04 1.0000000 4
## 1083 GO:0009942 3.107842e-04 0.9999803 6
## 1845 GO:0030488 3.134078e-04 0.9999528 9
## 1293 GO:0010305 3.670541e-04 0.9999189 12
## 357 GO:0006306 5.050144e-04 0.9999185 9
## 2758 GO:0048564 5.203432e-04 0.9999257 8
## 342 GO:0006281 5.519142e-04 0.9997526 31
## 1220 GO:0010190 5.714862e-04 0.9999670 5
## 1406 GO:0010583 6.182948e-04 0.9998525 12
## 1183 GO:0010109 6.783444e-04 0.9999431 6
## 992 GO:0009744 6.829633e-04 0.9997692 18
## 809 GO:0009088 7.057632e-04 0.9999403 6
## numInCat term ontology
## 949 185 chloroplast organization BP
## 882 213 response to light stimulus BP
## 1003 24 photosynthesis, light harvesting in photosystem I BP
## 3625 21 chloroplast rRNA processing BP
## 1556 63 photosynthesis BP
## 1560 42 chlorophyll biosynthetic process BP
## 1123 51 thylakoid membrane organization BP
## 1273 8 NADH dehydrogenase complex (plastoquinone) assembly BP
## 2333 11 plastid transcription BP
## 760 106 circadian rhythm BP
## 1006 18 photosynthetic electron transport in photosystem I BP
## 939 10 response to low light intensity stimulus BP
## 379 1468 regulation of transcription, DNA-templated BP
## 2486 20 protein targeting to chloroplast BP
## 876 349 response to cold BP
## 1258 10 chloroplast mRNA processing BP
## 938 46 response to high light intensity BP
## 1223 15 nonphotochemical quenching BP
## 1233 25 photosystem II assembly BP
## 940 41 response to absence of light BP
## 2487 28 protein import into chloroplast stroma BP
## 683 63 microtubule-based movement BP
## 948 13 plastid organization BP
## 1187 56 response to red light BP
## 1116 23 chloroplast fission BP
## 3225 11 cellular response to light stimulus BP
## 1002 21 photosynthetic electron transport chain BP
## 227 17 starch catabolic process BP
## 1906 14 chloroplast RNA processing BP
## 931 50 response to blue light BP
## 1590 7 carotene biosynthetic process BP
## 1117 7 amylopectin biosynthetic process BP
## 1015 468 embryo development ending in seed dormancy BP
## 69 26 Group II intron splicing BP
## 2722 140 leaf development BP
## 1004 10 photosynthesis, light harvesting in photosystem II BP
## 932 24 phototropism BP
## 146 6 rRNA 5'-end processing BP
## 1001 6 photosynthesis, light harvesting BP
## 1593 8 xanthophyll biosynthetic process BP
## 3607 22 chloroplast RNA modification BP
## 3608 4 chloroplast mRNA modification BP
## 1167 60 phloem or xylem histogenesis BP
## 1703 38 starch biosynthetic process BP
## 987 73 response to gibberellin BP
## 375 99 transcription, DNA-templated BP
## 238 15 fructose 6-phosphate metabolic process BP
## 1993 22 plastid translation BP
## 2411 4 RNA stabilization BP
## 1083 9 longitudinal axis specification BP
## 1845 20 tRNA methylation BP
## 1293 33 leaf vascular tissue pattern formation BP
## 357 20 DNA methylation BP
## 2758 18 photosystem I assembly BP
## 342 130 DNA repair BP
## 1220 8 cytochrome b6f complex assembly BP
## 1406 35 response to cyclopentenone BP
## 1183 10 regulation of photosynthesis BP
## 992 67 response to sucrose BP
## 809 10 threonine biosynthetic process BP
## over_represented_padjust
## 949 5.372569e-27
## 882 1.276386e-11
## 1003 3.223512e-11
## 3625 3.370762e-11
## 1556 1.591013e-09
## 1560 8.781955e-09
## 1123 8.142826e-08
## 1273 1.889667e-06
## 2333 4.868484e-06
## 760 4.868484e-06
## 1006 7.507505e-06
## 939 1.095835e-05
## 379 1.095835e-05
## 2486 2.429876e-05
## 876 2.963560e-05
## 1258 3.118698e-05
## 938 5.844619e-05
## 1223 1.575508e-04
## 1233 1.938647e-04
## 940 4.258646e-04
## 2487 5.026738e-04
## 683 7.951265e-04
## 948 7.951265e-04
## 1187 7.951265e-04
## 1116 9.479183e-04
## 3225 1.207912e-03
## 1002 1.658620e-03
## 227 1.800530e-03
## 1906 2.082647e-03
## 931 3.373289e-03
## 1590 4.188231e-03
## 1117 4.951329e-03
## 1015 5.259874e-03
## 69 5.508765e-03
## 2722 6.348064e-03
## 1004 7.769283e-03
## 932 1.056491e-02
## 146 1.091390e-02
## 1001 1.091390e-02
## 1593 1.091390e-02
## 3607 1.498762e-02
## 3608 1.498762e-02
## 1167 1.584437e-02
## 1703 1.584437e-02
## 987 1.827710e-02
## 375 2.053518e-02
## 238 2.281857e-02
## 1993 2.390907e-02
## 2411 2.390907e-02
## 1083 2.432905e-02
## 1845 2.432905e-02
## 1293 2.794552e-02
## 357 3.772362e-02
## 2758 3.814886e-02
## 342 3.972778e-02
## 1220 4.040203e-02
## 1406 4.294437e-02
## 1183 4.582799e-02
## 992 4.582799e-02
## 809 4.656861e-02do separate analysis for double coefficient type
for(n in 1:2) {
library(dplyr)
#genelist.up<-base::get(paste("../output/",DEG.objs.V1.5annotation.unique[n],sep="")) %>% rownames_to_column() %>% dplyr::filter(logFC>0) # does not work
genelist.all<-DEG.map[[n]]
# DEG.map[[n]] %>% dplyr::select("logFC.genotypenpr1.1")
# DEG.map[[n]] %>% dplyr::select(logFC.genotypenpr1.1)
#
# DEG.map[[n]] %>% dplyr::select(names(DEG.map[[n]])[2]) %>% filter(.>0)
# DEG.map[[n]] %>% dplyr::filter(names(DEG.map[[n]])[2]>0)
# DEG.map[[n]] %>% dplyr::filter(logFC.genotypenpr1.1>0) # does work
# DEG.map[[n]] %>% dplyr::filter("logFC.genotypenpr1.1">0)
# DEG.map[[n]] %>% dplyr::filter(rlang::UQ("logFC.genotypenpr1.1")>0)
# DEG.map[[n]] %>% dplyr::filter(!!"logFC.genotypenpr1.1">0)
# DEG.map[[n]] %>% dplyr::filter(!!names(DEG.map[[n]])[2]>0)
# DEG.map[[n]] %>% dplyr::filter((!!names(DEG.map[[n]])[2])==3.26)
# DEG.map[[n]] %>% dplyr::filter((!!names(DEG.map[[n]])[2])<0)
# DEG.map[[n]] %>% dplyr::filter((UQ(as.name(names(DEG.map[[n]])[2])))>0) # works! See https://stackoverflow.com/questions/27197617/filter-data-frame-by-character-column-name-in-dplyr
# DEG.map[[n]] %>% dplyr::filter((as.name(names(DEG.map[[n]])[2]))>0) # does not work
genelist.upup<-DEG.map[[n]] %>% dplyr::filter((UQ(as.name(names(DEG.map[[n]])[2])))>0) %>% dplyr::filter((UQ(as.name(names(DEG.map[[n]])[3])))>0)
genelist.updown<-DEG.map[[n]] %>% dplyr::filter((UQ(as.name(names(DEG.map[[n]])[2])))>0) %>% dplyr::filter((UQ(as.name(names(DEG.map[[n]])[3])))<0)
genelist.downup<-DEG.map[[n]] %>% dplyr::filter((UQ(as.name(names(DEG.map[[n]])[2])))<0) %>% dplyr::filter((UQ(as.name(names(DEG.map[[n]])[3])))>0)
genelist.downdown<- DEG.map[[n]] %>% dplyr::filter((UQ(as.name(names(DEG.map[[n]])[2])))<0) %>% dplyr::filter((UQ(as.name(names(DEG.map[[n]])[3])))<0)
GO.ORA.temp.all<-GOseq.Atgoslim.BP.list.ORA(str_remove(as_vector(genelist.all[,"genes"]),"\\.[[:digit:]]+"),custom.category.list=Atgoslim.TAIR.BP.list)
GO.ORA.temp.up_up<-GOseq.Atgoslim.BP.list.ORA(str_remove(as_vector(genelist.upup[,"genes"]),"\\.[[:digit:]]+"),custom.category.list=Atgoslim.TAIR.BP.list)
GO.ORA.temp.up_down<-GOseq.Atgoslim.BP.list.ORA(str_remove(as_vector(genelist.updown[,"genes"]),"\\.[[:digit:]]+"),custom.category.list=Atgoslim.TAIR.BP.list)
GO.ORA.temp.down_up<-GOseq.Atgoslim.BP.list.ORA(str_remove(as_vector(genelist.downup[,"genes"]),"\\.[[:digit:]]+"),custom.category.list=Atgoslim.TAIR.BP.list)
GO.ORA.temp.down_down<-GOseq.Atgoslim.BP.list.ORA(str_remove(as_vector(genelist.downdown[,"genes"]),"\\.[[:digit:]]+"),custom.category.list=Atgoslim.TAIR.BP.list)
# handling "no enriched GO"
# genelist.names<-c("GO.ORA.temp.up_down","GO.ORA.temp.down_up") # test
x<-list(GO.ORA.temp.all=GO.ORA.temp.all,
GO.ORA.temp.up_up=GO.ORA.temp.up_up,
GO.ORA.temp.up_down=GO.ORA.temp.up_down,
GO.ORA.temp.down_up=GO.ORA.temp.down_up,
GO.ORA.temp.down_down=GO.ORA.temp.down_down) # list
print(x[x=="no enriched GO"])
x<-x[!x=="no enriched GO"] # reove "no enriched GO" result
## add sample info and FC info and save GO.ORA result
for (i in 1:length(x)) {
GO.ORA.result<-x[[i]] %>% mutate(FC = gsub("(GO.ORA.temp.)(.)","\\2",names(x)[i]),sample=DEG.objs[n])
save(GO.ORA.result,file=file.path("output",paste(gsub(".csv","",DEG.objs2[n]),gsub("(GO.ORA.temp.)(.)","\\2",names(x)[i]),"enrich.Rdata",sep=".")))
rm(GO.ORA.result)
}
}## Warning in pcls(G): initial point very close to some inequality constraints## Using manually entered categories.## Calculating the p-values...## 'select()' returned 1:1 mapping between keys and columns## [1] "enriched.GO is"
## category over_represented_pvalue under_represented_pvalue numDEInCat
## 3214 GO:0071456 6.866937e-82 1.0000000 187
## 2314 GO:0042742 7.898946e-35 1.0000000 200
## 1227 GO:0010200 4.003411e-29 1.0000000 91
## 444 GO:0006468 1.636305e-26 1.0000000 363
## 919 GO:0009617 5.830579e-22 1.0000000 75
## 3367 GO:0080167 7.029884e-22 1.0000000 78
## 997 GO:0009751 1.988635e-19 1.0000000 67
## 945 GO:0009651 9.065322e-18 1.0000000 163
## 882 GO:0009416 2.160288e-17 1.0000000 103
## 875 GO:0009408 3.071216e-17 1.0000000 87
## 737 GO:0007165 6.380586e-17 1.0000000 165
## 2824 GO:0050832 1.224297e-16 1.0000000 103
## 880 GO:0009414 3.481991e-16 1.0000000 133
## 915 GO:0009611 2.417818e-15 1.0000000 98
## 876 GO:0009409 6.627074e-15 1.0000000 140
## 665 GO:0006979 2.387508e-12 1.0000000 117
## 923 GO:0009626 5.733685e-12 1.0000000 33
## 379 GO:0006355 8.467621e-12 1.0000000 418
## 985 GO:0009737 9.189281e-12 1.0000000 155
## 920 GO:0009620 8.178923e-11 1.0000000 51
## 1003 GO:0009768 9.484722e-11 1.0000000 20
## 658 GO:0006952 1.368428e-10 1.0000000 77
## 2299 GO:0042542 8.704672e-10 1.0000000 35
## 924 GO:0009627 4.294687e-09 1.0000000 31
## 199 GO:0002237 6.232604e-09 1.0000000 24
## 1187 GO:0010114 1.356273e-08 1.0000000 32
## 3507 GO:0098542 1.654445e-08 1.0000000 27
## 760 GO:0007623 3.092534e-08 1.0000000 50
## 981 GO:0009733 8.570350e-08 1.0000000 85
## 1560 GO:0015995 1.007216e-07 1.0000000 26
## 661 GO:0006970 1.313257e-07 0.9999999 62
## 938 GO:0009644 1.355660e-07 1.0000000 26
## 233 GO:0005992 1.538097e-07 1.0000000 17
## 2654 GO:0046777 1.837007e-07 0.9999999 77
## 3601 GO:1900426 2.482052e-07 1.0000000 19
## 1643 GO:0016998 3.941070e-07 1.0000000 13
## 738 GO:0007166 1.345690e-06 0.9999997 26
## 1222 GO:0010193 1.846904e-06 0.9999997 20
## 2646 GO:0046686 1.977003e-06 0.9999993 107
## 992 GO:0009744 2.084806e-06 0.9999994 32
## 931 GO:0009637 2.102416e-06 0.9999995 27
## 160 GO:0001666 2.932318e-06 0.9999994 23
## 201 GO:0002239 3.933864e-06 0.9999995 16
## 998 GO:0009753 4.336867e-06 0.9999983 47
## 200 GO:0002238 4.455554e-06 0.9999996 13
## 3881 GO:2000022 4.464626e-06 0.9999990 23
## 1899 GO:0031347 5.383413e-06 0.9999986 27
## 1240 GO:0010218 6.992208e-06 0.9999984 23
## 986 GO:0009738 8.071132e-06 0.9999970 41
## 2497 GO:0045087 1.040445e-05 0.9999967 31
## 922 GO:0009625 1.091963e-05 0.9999979 19
## 2869 GO:0051259 1.240682e-05 0.9999988 11
## 987 GO:0009739 1.397674e-05 0.9999954 32
## 197 GO:0002229 1.991412e-05 0.9999932 34
## 2561 GO:0045893 2.278827e-05 0.9999874 92
## 2207 GO:0035556 2.643298e-05 0.9999867 69
## 1628 GO:0016567 2.899255e-05 0.9999808 175
## 1611 GO:0016310 3.190967e-05 0.9999823 93
## 921 GO:0009624 3.515701e-05 0.9999868 36
## 940 GO:0009646 3.677165e-05 0.9999909 21
## 1246 GO:0010224 3.697906e-05 0.9999877 30
## 441 GO:0006457 3.959960e-05 0.9999820 50
## 3921 GO:2000280 4.348249e-05 0.9999875 23
## 3088 GO:0070370 4.955656e-05 0.9999972 9
## 934 GO:0009640 5.007753e-05 0.9999838 28
## 2560 GO:0045892 5.382916e-05 0.9999732 60
## 3348 GO:0080142 7.618140e-05 0.9999938 10
## 3226 GO:0071483 9.880168e-05 1.0000000 6
## 2366 GO:0043069 1.104694e-04 0.9999870 11
## 2757 GO:0048544 1.207463e-04 0.9999686 20
## 683 GO:0007018 1.329515e-04 0.9999533 29
## 957 GO:0009682 1.407146e-04 0.9999770 13
## 1193 GO:0010120 1.973179e-04 0.9999830 9
## 2141 GO:0034605 2.303414e-04 0.9999208 25
## 1000 GO:0009759 2.465032e-04 0.9999909 7
## 995 GO:0009749 2.960107e-04 0.9999063 22
## 949 GO:0009658 3.057114e-04 0.9998331 63
## 2512 GO:0045332 3.074021e-04 0.9999716 9
## 2144 GO:0034620 3.161043e-04 0.9999540 11
## 3422 GO:0090333 3.867742e-04 0.9998998 17
## 3164 GO:0071215 5.173413e-04 0.9998582 17
## 1425 GO:0010629 5.176749e-04 0.9999023 12
## 659 GO:0006955 5.205667e-04 0.9999401 9
## 3901 GO:2000067 5.401621e-04 0.9998768 13
## 2722 GO:0048366 5.810771e-04 0.9997019 48
## 1188 GO:0010115 5.861408e-04 0.9999603 7
## 971 GO:0009699 6.243803e-04 0.9999408 8
## 192 GO:0002213 6.551291e-04 0.9998585 13
## 2333 GO:0042793 6.811799e-04 0.9999344 8
## 2952 GO:0051865 6.874629e-04 0.9998824 11
## 269 GO:0006094 7.074251e-04 0.9998499 13
## 1212 GO:0010167 7.222927e-04 0.9998756 11
## 2283 GO:0042344 7.246484e-04 1.0000000 5
## 1175 GO:0010099 7.649301e-04 0.9998902 9
## 2928 GO:0051607 8.255408e-04 0.9997185 21
## 605 GO:0006816 8.578817e-04 0.9997557 17
## 628 GO:0006874 8.989047e-04 0.9997296 18
## 3210 GO:0071446 9.314771e-04 0.9999310 7
## 989 GO:0009741 9.671433e-04 0.9997227 16
## 2938 GO:0051707 9.875533e-04 0.9997758 13
## 206 GO:0002758 1.028506e-03 0.9997653 13
## 333 GO:0006267 1.130641e-03 0.9999505 6
## 2819 GO:0050801 1.138254e-03 0.9998634 8
## 932 GO:0009638 1.187348e-03 0.9997242 13
## 794 GO:0009051 1.204628e-03 0.9998263 9
## 2159 GO:0034758 1.257269e-03 1.0000000 3
## 939 GO:0009645 1.304618e-03 0.9998729 7
## 1288 GO:0010286 1.339388e-03 0.9995148 21
## 196 GO:0002221 1.373614e-03 0.9997977 9
## 1165 GO:0010082 1.377524e-03 0.9995819 16
## 1280 GO:0010268 1.382294e-03 0.9996452 14
## 3868 GO:1990641 1.397819e-03 0.9998101 8
## 2195 GO:0035435 1.403742e-03 0.9998253 8
## numInCat
## 3214 238
## 2314 391
## 1227 138
## 444 928
## 919 118
## 3367 128
## 997 108
## 945 395
## 882 213
## 875 169
## 737 411
## 2824 218
## 880 318
## 915 209
## 876 349
## 665 301
## 923 46
## 379 1468
## 985 423
## 920 99
## 1003 24
## 658 192
## 2299 62
## 924 54
## 199 34
## 1187 56
## 3507 44
## 760 106
## 981 251
## 1560 42
## 661 152
## 938 46
## 233 22
## 2654 188
## 3601 27
## 1643 16
## 738 46
## 1222 34
## 2646 312
## 992 67
## 931 50
## 160 40
## 201 24
## 998 113
## 200 17
## 3881 42
## 1899 53
## 1240 43
## 986 99
## 2497 68
## 922 33
## 2869 17
## 987 73
## 197 74
## 2561 284
## 2207 185
## 1628 608
## 1611 273
## 921 86
## 940 41
## 1246 67
## 441 138
## 3921 54
## 3088 11
## 934 62
## 2560 173
## 3348 13
## 3226 6
## 2366 16
## 2757 38
## 683 63
## 957 21
## 1193 12
## 2141 59
## 1000 8
## 995 48
## 949 185
## 2512 12
## 2144 17
## 3422 33
## 3164 35
## 1425 21
## 659 13
## 3901 28
## 2722 140
## 1188 9
## 971 11
## 192 24
## 2333 11
## 2952 18
## 269 23
## 1212 18
## 2283 5
## 1175 15
## 2928 48
## 605 34
## 628 37
## 3210 9
## 989 35
## 2938 24
## 206 24
## 333 7
## 2819 12
## 932 24
## 794 14
## 2159 3
## 939 10
## 1288 51
## 196 14
## 1165 37
## 1280 27
## 3868 13
## 2195 12
## term
## 3214 cellular response to hypoxia
## 2314 defense response to bacterium
## 1227 response to chitin
## 444 protein phosphorylation
## 919 response to bacterium
## 3367 response to karrikin
## 997 response to salicylic acid
## 945 response to salt stress
## 882 response to light stimulus
## 875 response to heat
## 737 signal transduction
## 2824 defense response to fungus
## 880 response to water deprivation
## 915 response to wounding
## 876 response to cold
## 665 response to oxidative stress
## 923 plant-type hypersensitive response
## 379 regulation of transcription, DNA-templated
## 985 response to abscisic acid
## 920 response to fungus
## 1003 photosynthesis, light harvesting in photosystem I
## 658 defense response
## 2299 response to hydrogen peroxide
## 924 systemic acquired resistance
## 199 response to molecule of bacterial origin
## 1187 response to red light
## 3507 defense response to other organism
## 760 circadian rhythm
## 981 response to auxin
## 1560 chlorophyll biosynthetic process
## 661 response to osmotic stress
## 938 response to high light intensity
## 233 trehalose biosynthetic process
## 2654 protein autophosphorylation
## 3601 positive regulation of defense response to bacterium
## 1643 cell wall macromolecule catabolic process
## 738 cell surface receptor signaling pathway
## 1222 response to ozone
## 2646 response to cadmium ion
## 992 response to sucrose
## 931 response to blue light
## 160 response to hypoxia
## 201 response to oomycetes
## 998 response to jasmonic acid
## 200 response to molecule of fungal origin
## 3881 regulation of jasmonic acid mediated signaling pathway
## 1899 regulation of defense response
## 1240 response to far red light
## 986 abscisic acid-activated signaling pathway
## 2497 innate immune response
## 922 response to insect
## 2869 protein complex oligomerization
## 987 response to gibberellin
## 197 defense response to oomycetes
## 2561 positive regulation of transcription, DNA-templated
## 2207 intracellular signal transduction
## 1628 protein ubiquitination
## 1611 phosphorylation
## 921 response to nematode
## 940 response to absence of light
## 1246 response to UV-B
## 441 protein folding
## 3921 regulation of root development
## 3088 cellular heat acclimation
## 934 photomorphogenesis
## 2560 negative regulation of transcription, DNA-templated
## 3348 regulation of salicylic acid biosynthetic process
## 3226 cellular response to blue light
## 2366 negative regulation of programmed cell death
## 2757 recognition of pollen
## 683 microtubule-based movement
## 957 induced systemic resistance
## 1193 camalexin biosynthetic process
## 2141 cellular response to heat
## 1000 indole glucosinolate biosynthetic process
## 995 response to glucose
## 949 chloroplast organization
## 2512 phospholipid translocation
## 2144 cellular response to unfolded protein
## 3422 regulation of stomatal closure
## 3164 cellular response to abscisic acid stimulus
## 1425 negative regulation of gene expression
## 659 immune response
## 3901 regulation of root morphogenesis
## 2722 leaf development
## 1188 regulation of abscisic acid biosynthetic process
## 971 phenylpropanoid biosynthetic process
## 192 defense response to insect
## 2333 plastid transcription
## 2952 protein autoubiquitination
## 269 gluconeogenesis
## 1212 response to nitrate
## 2283 indole glucosinolate catabolic process
## 1175 regulation of photomorphogenesis
## 2928 defense response to virus
## 605 calcium ion transport
## 628 cellular calcium ion homeostasis
## 3210 cellular response to salicylic acid stimulus
## 989 response to brassinosteroid
## 2938 response to other organism
## 206 innate immune response-activating signal transduction
## 333 pre-replicative complex assembly involved in nuclear cell cycle DNA replication
## 2819 ion homeostasis
## 932 phototropism
## 794 pentose-phosphate shunt, oxidative branch
## 2159 positive regulation of iron ion transport
## 939 response to low light intensity stimulus
## 1288 heat acclimation
## 196 pattern recognition receptor signaling pathway
## 1165 regulation of root meristem growth
## 1280 brassinosteroid homeostasis
## 3868 response to iron ion starvation
## 2195 phosphate ion transmembrane transport
## ontology over_represented_padjust
## 3214 BP 2.718620e-78
## 2314 BP 1.563596e-31
## 1227 BP 5.283168e-26
## 444 BP 1.619533e-23
## 919 BP 4.616653e-19
## 3367 BP 4.638552e-19
## 997 BP 1.124715e-16
## 945 BP 4.486201e-15
## 882 BP 9.502865e-15
## 875 BP 1.215894e-14
## 737 BP 2.296431e-14
## 2824 BP 4.039160e-14
## 880 BP 1.060400e-13
## 915 BP 6.837243e-13
## 876 BP 1.749106e-12
## 665 BP 5.907590e-10
## 923 BP 1.335274e-09
## 379 BP 1.862406e-09
## 985 BP 1.914756e-09
## 920 BP 1.619018e-08
## 1003 BP 1.788096e-08
## 658 BP 2.462548e-08
## 2299 BP 1.498339e-07
## 924 BP 7.084444e-07
## 199 BP 9.869952e-07
## 1187 BP 2.065186e-06
## 3507 BP 2.425906e-06
## 760 BP 4.372623e-06
## 981 BP 1.170001e-05
## 1560 BP 1.329189e-05
## 661 BP 1.677156e-05
## 938 BP 1.677205e-05
## 233 BP 1.845250e-05
## 2654 BP 2.139033e-05
## 3601 BP 2.807555e-05
## 1643 BP 4.334082e-05
## 738 BP 1.439889e-04
## 1222 BP 1.924183e-04
## 2646 BP 2.006912e-04
## 992 BP 2.030114e-04
## 931 BP 2.030114e-04
## 160 BP 2.764059e-04
## 201 BP 3.621899e-04
## 998 BP 3.842490e-04
## 200 BP 3.842490e-04
## 3881 BP 3.842490e-04
## 1899 BP 4.534667e-04
## 1240 BP 5.767115e-04
## 986 BP 6.521145e-04
## 2497 BP 8.238244e-04
## 922 BP 8.476632e-04
## 2869 BP 9.445884e-04
## 987 BP 1.044036e-03
## 197 BP 1.460000e-03
## 2561 BP 1.640341e-03
## 2207 BP 1.868717e-03
## 1628 BP 2.013710e-03
## 1611 BP 2.178110e-03
## 921 BP 2.359095e-03
## 940 BP 2.400002e-03
## 1246 BP 2.400002e-03
## 441 BP 2.528626e-03
## 3921 BP 2.732495e-03
## 3088 BP 3.050107e-03
## 934 BP 3.050107e-03
## 2560 BP 3.228934e-03
## 3348 BP 4.501525e-03
## 3226 BP 5.752292e-03
## 2366 BP 6.338382e-03
## 2757 BP 6.829065e-03
## 683 BP 7.413448e-03
## 957 BP 7.737351e-03
## 1193 BP 1.070112e-02
## 2141 BP 1.232327e-02
## 1000 BP 1.301208e-02
## 995 BP 1.541982e-02
## 949 BP 1.560263e-02
## 2512 BP 1.560263e-02
## 2144 BP 1.584123e-02
## 3422 BP 1.914049e-02
## 3164 BP 2.483040e-02
## 1425 BP 2.483040e-02
## 659 BP 2.483040e-02
## 3901 BP 2.545835e-02
## 2722 BP 2.698292e-02
## 1188 BP 2.698292e-02
## 971 BP 2.841289e-02
## 192 BP 2.947336e-02
## 2333 BP 3.024073e-02
## 2952 BP 3.024073e-02
## 269 BP 3.077688e-02
## 1212 BP 3.084820e-02
## 2283 BP 3.084820e-02
## 1175 BP 3.221658e-02
## 2928 BP 3.440333e-02
## 605 BP 3.537868e-02
## 628 BP 3.668828e-02
## 3210 BP 3.762977e-02
## 989 BP 3.867596e-02
## 2938 BP 3.909724e-02
## 206 BP 4.031538e-02
## 333 BP 4.375095e-02
## 2819 BP 4.375095e-02
## 932 BP 4.519913e-02
## 794 BP 4.542020e-02
## 2159 BP 4.695782e-02
## 939 BP 4.827088e-02
## 1288 BP 4.909848e-02
## 196 BP 4.918068e-02
## 1165 BP 4.918068e-02
## 1280 BP 4.918068e-02
## 3868 BP 4.918068e-02
## 2195 BP 4.918068e-02## Warning in pcls(G): initial point very close to some inequality constraints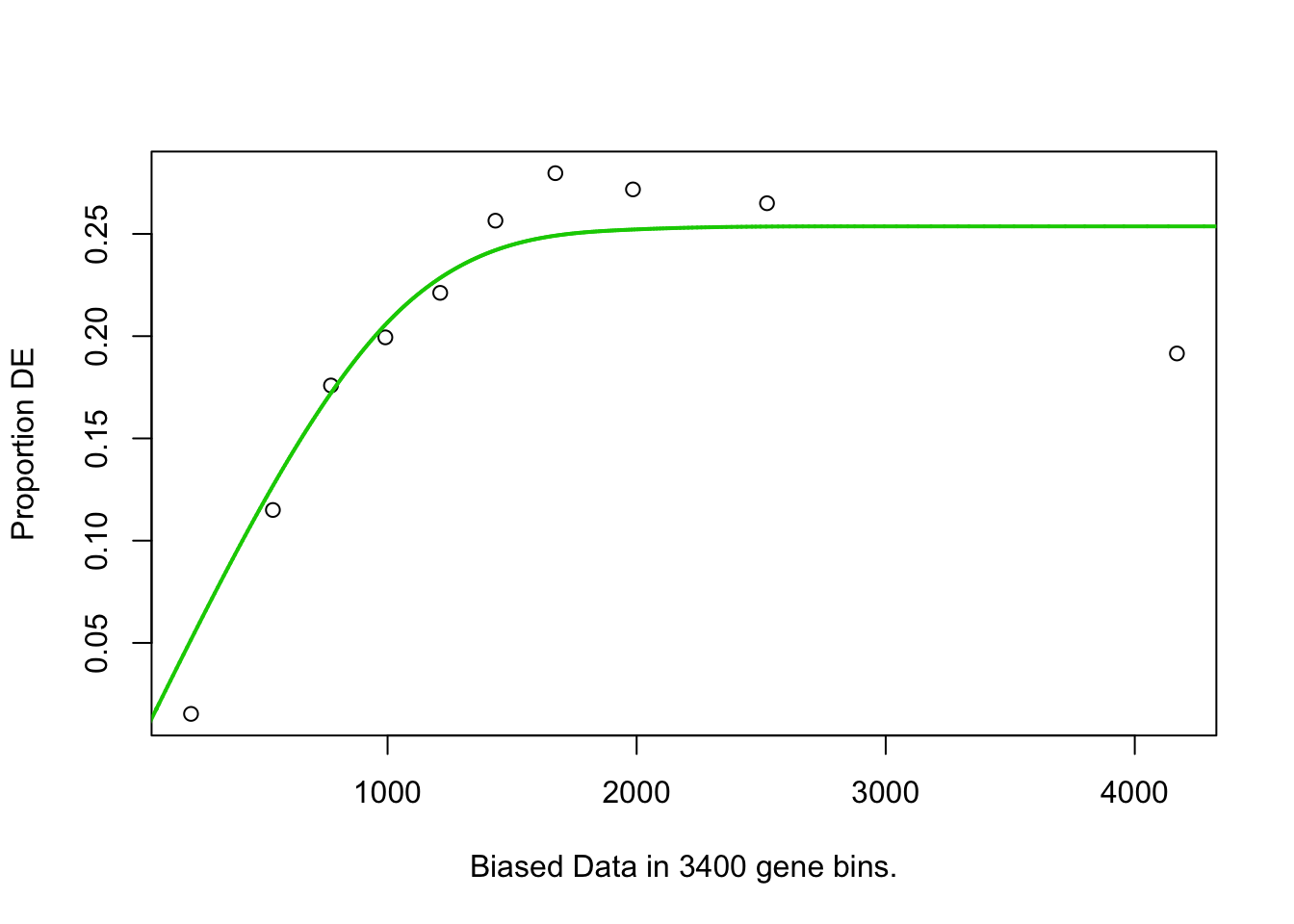
## Using manually entered categories.## Calculating the p-values...## 'select()' returned 1:1 mapping between keys and columns## [1] "enriched.GO is"
## category over_represented_pvalue under_represented_pvalue numDEInCat
## 1003 GO:0009768 6.432748e-21 1.0000000 18
## 882 GO:0009416 8.593302e-15 1.0000000 39
## 1560 GO:0015995 8.393074e-13 1.0000000 17
## 683 GO:0007018 3.249882e-11 1.0000000 19
## 939 GO:0009645 2.108753e-08 1.0000000 7
## 876 GO:0009409 2.758485e-08 1.0000000 39
## 949 GO:0009658 3.839056e-08 1.0000000 27
## 1556 GO:0015979 8.495700e-08 1.0000000 14
## 1004 GO:0009769 1.621533e-07 1.0000000 6
## 938 GO:0009644 1.461428e-06 0.9999998 11
## 760 GO:0007623 3.433846e-06 0.9999992 17
## 1207 GO:0010158 7.081939e-06 0.9999998 5
## 1817 GO:0030104 1.638744e-05 0.9999995 5
## 1187 GO:0010114 1.899371e-05 0.9999969 11
## 587 GO:0006782 3.089870e-05 0.9999981 6
## 1123 GO:0010027 4.728019e-05 0.9999924 10
## 2333 GO:0042793 6.175741e-05 0.9999972 5
## 1240 GO:0010218 6.432789e-05 0.9999906 9
## 928 GO:0009631 9.802492e-05 0.9999825 10
## 133 GO:0000910 1.212143e-04 1.0000000 3
## 614 GO:0006833 1.507631e-04 0.9999865 6
## 3751 GO:1903428 1.958414e-04 0.9999981 3
## 375 GO:0006351 2.449631e-04 0.9999365 13
## 3607 GO:1900865 2.704392e-04 0.9999726 6
## 2722 GO:0048366 2.721632e-04 0.9999160 16
## 851 GO:0009263 3.221376e-04 0.9999964 3
## numInCat
## 1003 24
## 882 213
## 1560 42
## 683 63
## 939 10
## 876 349
## 949 185
## 1556 63
## 1004 10
## 938 46
## 760 106
## 1207 8
## 1817 9
## 1187 56
## 587 15
## 1123 51
## 2333 11
## 1240 43
## 928 59
## 133 3
## 614 21
## 3751 4
## 375 99
## 3607 22
## 2722 140
## 851 4
## term
## 1003 photosynthesis, light harvesting in photosystem I
## 882 response to light stimulus
## 1560 chlorophyll biosynthetic process
## 683 microtubule-based movement
## 939 response to low light intensity stimulus
## 876 response to cold
## 949 chloroplast organization
## 1556 photosynthesis
## 1004 photosynthesis, light harvesting in photosystem II
## 938 response to high light intensity
## 760 circadian rhythm
## 1207 abaxial cell fate specification
## 1817 water homeostasis
## 1187 response to red light
## 587 protoporphyrinogen IX biosynthetic process
## 1123 thylakoid membrane organization
## 2333 plastid transcription
## 1240 response to far red light
## 928 cold acclimation
## 133 cytokinesis
## 614 water transport
## 3751 positive regulation of reactive oxygen species biosynthetic process
## 375 transcription, DNA-templated
## 3607 chloroplast RNA modification
## 2722 leaf development
## 851 deoxyribonucleotide biosynthetic process
## ontology over_represented_padjust
## 1003 BP 2.546725e-17
## 882 BP 1.701044e-11
## 1560 BP 1.107606e-09
## 683 BP 3.216571e-08
## 939 BP 1.669710e-05
## 876 BP 1.820140e-05
## 949 BP 2.171260e-05
## 1556 BP 4.204310e-05
## 1004 BP 7.132944e-05
## 938 BP 5.785791e-04
## 760 BP 1.235872e-03
## 1207 BP 2.336450e-03
## 1817 BP 4.990607e-03
## 1187 BP 5.371150e-03
## 587 BP 8.155197e-03
## 1123 BP 1.169889e-02
## 2333 BP 1.414856e-02
## 1240 BP 1.414856e-02
## 928 BP 2.042530e-02
## 133 BP 2.399436e-02
## 614 BP 2.842244e-02
## 3751 BP 3.524255e-02
## 375 BP 4.216560e-02
## 3607 BP 4.309977e-02
## 2722 BP 4.309977e-02
## 851 BP 4.905164e-02## Warning in pcls(G): initial point very close to some inequality constraints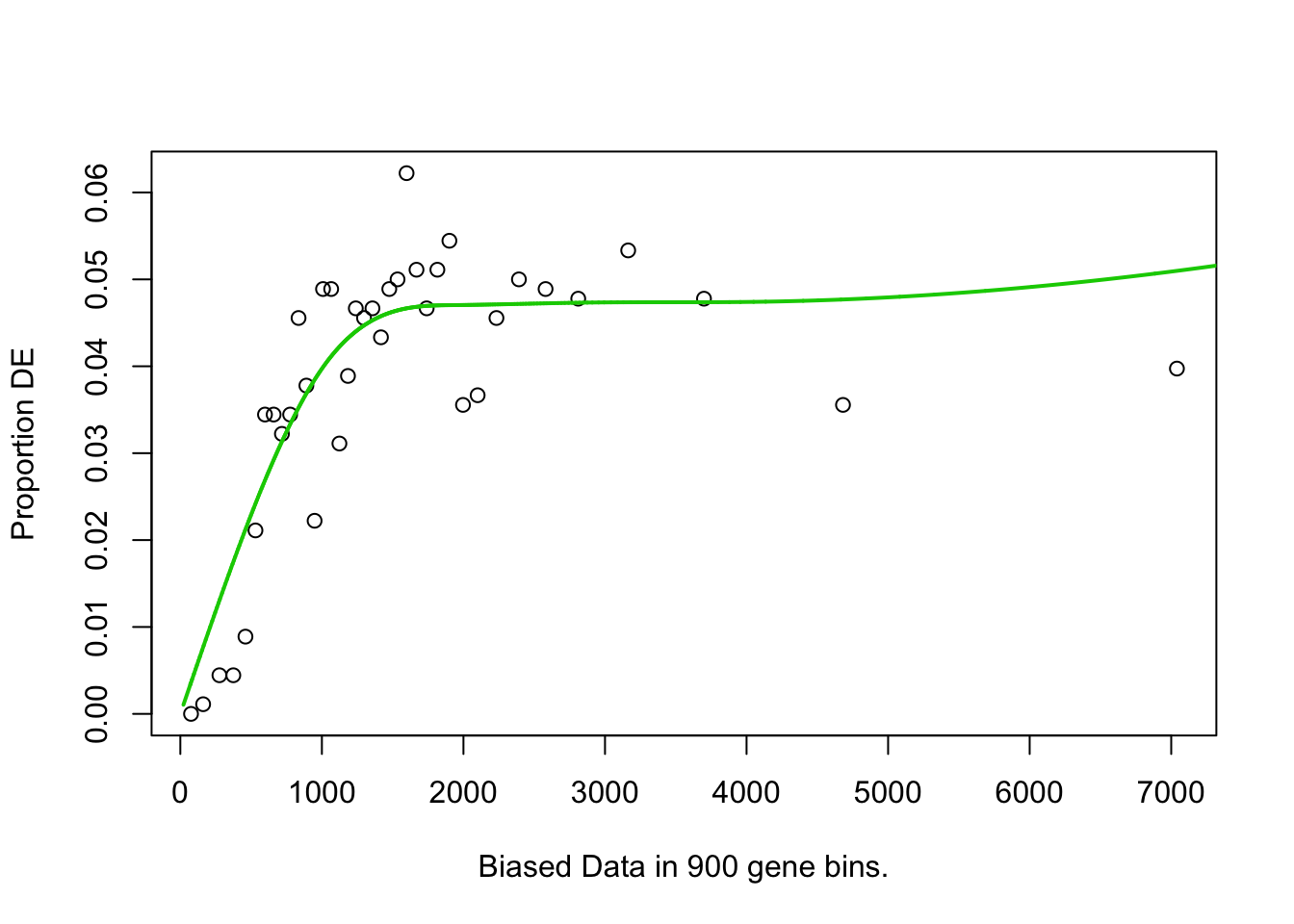
## Using manually entered categories.## Calculating the p-values...## 'select()' returned 1:1 mapping between keys and columns## [1] "enriched.GO is"
## category over_represented_pvalue under_represented_pvalue numDEInCat
## 945 GO:0009651 6.800812e-12 1.0000000 52
## 2869 GO:0051259 7.151062e-09 1.0000000 8
## 875 GO:0009408 1.126646e-08 1.0000000 27
## 2299 GO:0042542 1.713297e-07 1.0000000 14
## 333 GO:0006267 2.199669e-07 1.0000000 6
## 334 GO:0006268 2.922316e-07 1.0000000 9
## 1246 GO:0010224 2.947907e-07 1.0000000 15
## 441 GO:0006457 5.575957e-07 0.9999999 21
## 3367 GO:0080167 6.360004e-07 0.9999999 20
## 1643 GO:0016998 8.667331e-07 1.0000000 7
## 3226 GO:0071483 8.926728e-07 1.0000000 5
## 998 GO:0009753 1.016816e-06 0.9999998 19
## 2159 GO:0034758 4.836935e-06 1.0000000 3
## 915 GO:0009611 1.420503e-05 0.9999953 25
## 121 GO:0000727 1.625592e-05 0.9999992 6
## 985 GO:0009737 1.819454e-05 0.9999920 40
## 378 GO:0006354 3.550457e-05 0.9999996 4
## 880 GO:0009414 3.699313e-05 0.9999852 31
## 2561 GO:0045893 5.475384e-05 0.9999787 28
## 1363 GO:0010438 1.052790e-04 0.9999965 4
## 1023 GO:0009809 1.191007e-04 0.9999811 9
## 2318 GO:0042754 1.407189e-04 1.0000000 3
## 1252 GO:0010230 1.993693e-04 0.9999930 4
## 813 GO:0009094 2.014812e-04 0.9999872 5
## numInCat
## 945 395
## 2869 17
## 875 169
## 2299 62
## 333 7
## 334 19
## 1246 67
## 441 138
## 3367 128
## 1643 16
## 3226 6
## 998 113
## 2159 3
## 915 209
## 121 12
## 985 423
## 378 5
## 880 318
## 2561 284
## 1363 10
## 1023 42
## 2318 3
## 1252 8
## 813 13
## term
## 945 response to salt stress
## 2869 protein complex oligomerization
## 875 response to heat
## 2299 response to hydrogen peroxide
## 333 pre-replicative complex assembly involved in nuclear cell cycle DNA replication
## 334 DNA unwinding involved in DNA replication
## 1246 response to UV-B
## 441 protein folding
## 3367 response to karrikin
## 1643 cell wall macromolecule catabolic process
## 3226 cellular response to blue light
## 998 response to jasmonic acid
## 2159 positive regulation of iron ion transport
## 915 response to wounding
## 121 double-strand break repair via break-induced replication
## 985 response to abscisic acid
## 378 DNA-templated transcription, elongation
## 880 response to water deprivation
## 2561 positive regulation of transcription, DNA-templated
## 1363 cellular response to sulfur starvation
## 1023 lignin biosynthetic process
## 2318 negative regulation of circadian rhythm
## 1252 alternative respiration
## 813 L-phenylalanine biosynthetic process
## ontology over_represented_padjust
## 945 BP 2.692442e-08
## 2869 BP 1.415553e-05
## 875 BP 1.486797e-05
## 2299 BP 1.667252e-04
## 333 BP 1.667252e-04
## 334 BP 1.667252e-04
## 1246 BP 1.667252e-04
## 441 BP 2.759402e-04
## 3367 BP 2.797695e-04
## 1643 BP 3.212810e-04
## 3226 BP 3.212810e-04
## 998 BP 3.354646e-04
## 2159 BP 1.473033e-03
## 915 BP 4.016981e-03
## 121 BP 4.290479e-03
## 985 BP 4.502013e-03
## 378 BP 8.136435e-03
## 880 BP 8.136435e-03
## 2561 BP 1.140897e-02
## 1363 BP 2.083998e-02
## 1023 BP 2.245332e-02
## 2318 BP 2.532300e-02
## 1252 BP 3.323601e-02
## 813 BP 3.323601e-02## Warning in pcls(G): initial point very close to some inequality constraints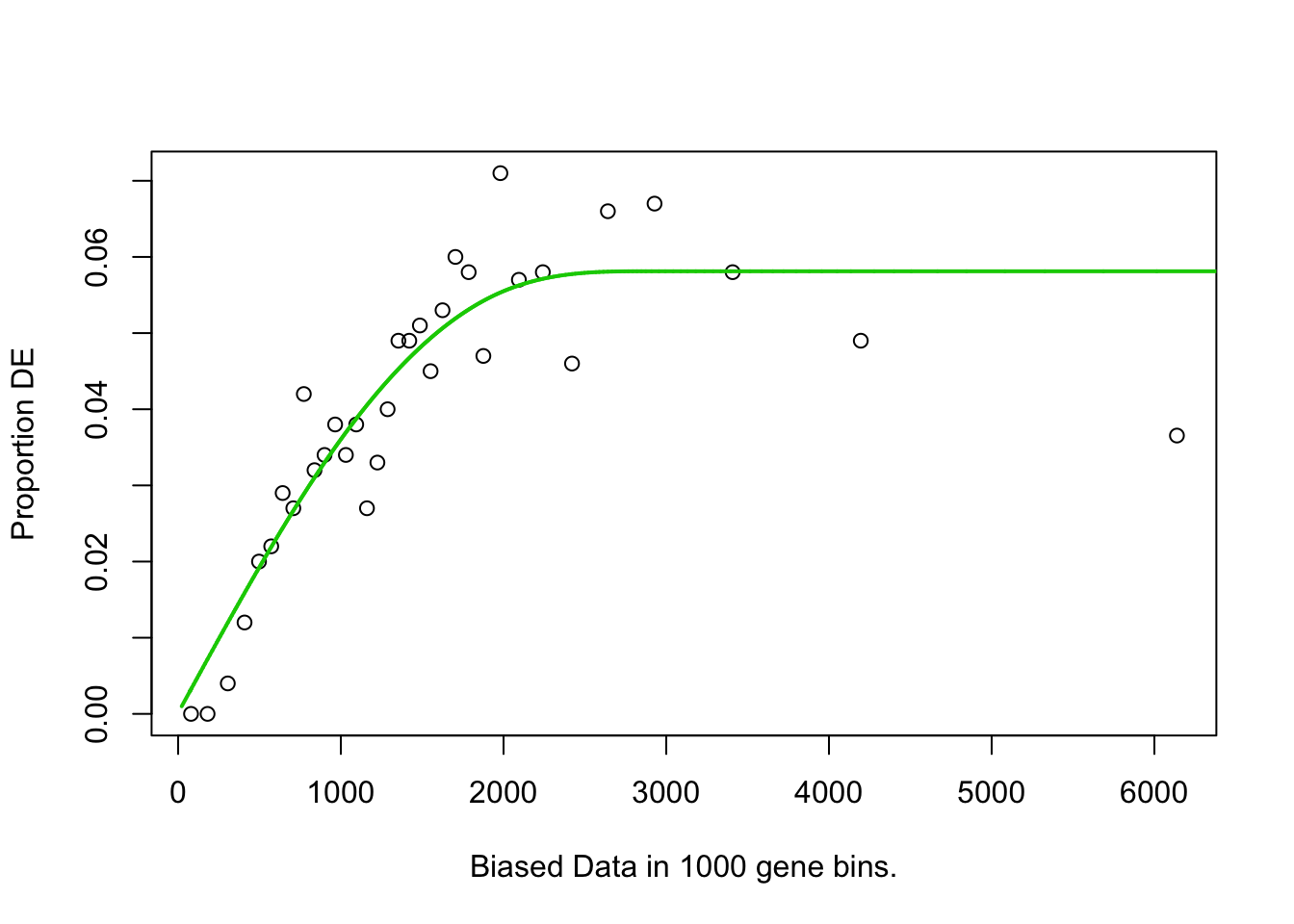
## Using manually entered categories.## Calculating the p-values...## 'select()' returned 1:1 mapping between keys and columns## [1] "enriched.GO is"
## category over_represented_pvalue under_represented_pvalue numDEInCat
## 3214 GO:0071456 2.796477e-53 1.0000000 98
## 379 GO:0006355 4.338524e-15 1.0000000 177
## 1227 GO:0010200 4.732198e-12 1.0000000 35
## 940 GO:0009646 4.126635e-11 1.0000000 18
## 981 GO:0009733 2.886609e-10 1.0000000 44
## 160 GO:0001666 6.122972e-09 1.0000000 16
## 1628 GO:0016567 2.898539e-08 1.0000000 78
## 931 GO:0009637 2.278063e-07 1.0000000 16
## 2314 GO:0042742 8.482207e-07 0.9999997 55
## 997 GO:0009751 8.683719e-07 0.9999998 23
## 882 GO:0009416 1.694619e-06 0.9999994 35
## 798 GO:0009061 1.951404e-06 1.0000000 6
## 923 GO:0009626 3.326760e-06 0.9999994 14
## 880 GO:0009414 5.610322e-06 0.9999976 44
## 989 GO:0009741 9.188483e-06 0.9999987 11
## 920 GO:0009620 1.102265e-05 0.9999970 20
## 992 GO:0009744 3.410390e-05 0.9999921 15
## 1425 GO:0010629 4.565716e-05 0.9999952 8
## 3367 GO:0080167 4.839729e-05 0.9999843 22
## 2366 GO:0043069 5.792747e-05 0.9999951 7
## 876 GO:0009409 7.299621e-05 0.9999648 44
## 987 GO:0009739 8.258091e-05 0.9999792 15
## 3632 GO:1901371 1.019753e-04 0.9999897 7
## 2753 GO:0048527 1.317688e-04 0.9999717 12
## 2283 GO:0042344 1.380116e-04 0.9999978 4
## 3363 GO:0080163 1.407585e-04 0.9999885 6
## 1212 GO:0010167 1.790356e-04 0.9999809 7
## 1059 GO:0009873 2.325999e-04 0.9999507 11
## 3314 GO:0080037 3.362756e-04 0.9999899 4
## numInCat term
## 3214 238 cellular response to hypoxia
## 379 1468 regulation of transcription, DNA-templated
## 1227 138 response to chitin
## 940 41 response to absence of light
## 981 251 response to auxin
## 160 40 response to hypoxia
## 1628 608 protein ubiquitination
## 931 50 response to blue light
## 2314 391 defense response to bacterium
## 997 108 response to salicylic acid
## 882 213 response to light stimulus
## 798 9 anaerobic respiration
## 923 46 plant-type hypersensitive response
## 880 318 response to water deprivation
## 989 35 response to brassinosteroid
## 920 99 response to fungus
## 992 67 response to sucrose
## 1425 21 negative regulation of gene expression
## 3367 128 response to karrikin
## 2366 16 negative regulation of programmed cell death
## 876 349 response to cold
## 987 73 response to gibberellin
## 3632 20 regulation of leaf morphogenesis
## 2753 53 lateral root development
## 2283 5 indole glucosinolate catabolic process
## 3363 16 regulation of protein serine/threonine phosphatase activity
## 1212 18 response to nitrate
## 1059 49 ethylene-activated signaling pathway
## 3314 6 negative regulation of cytokinin-activated signaling pathway
## ontology over_represented_padjust
## 3214 BP 1.107125e-49
## 379 BP 8.588109e-12
## 1227 BP 6.244924e-09
## 940 BP 4.084337e-08
## 981 BP 2.285617e-07
## 160 BP 4.040141e-06
## 1628 BP 1.639331e-05
## 931 BP 1.127356e-04
## 2314 BP 3.437884e-04
## 997 BP 3.437884e-04
## 882 BP 6.099088e-04
## 798 BP 6.438008e-04
## 923 BP 1.013126e-03
## 880 BP 1.586519e-03
## 989 BP 2.425147e-03
## 920 BP 2.727418e-03
## 992 BP 7.942196e-03
## 1425 BP 1.004204e-02
## 3367 BP 1.008447e-02
## 2366 BP 1.146674e-02
## 876 BP 1.376152e-02
## 987 BP 1.486081e-02
## 3632 BP 1.755306e-02
## 2753 BP 2.143320e-02
## 2283 BP 2.143320e-02
## 3363 BP 2.143320e-02
## 1212 BP 2.625193e-02
## 1059 BP 3.288796e-02
## 3314 BP 4.590742e-02## Warning in pcls(G): initial point very close to some inequality constraints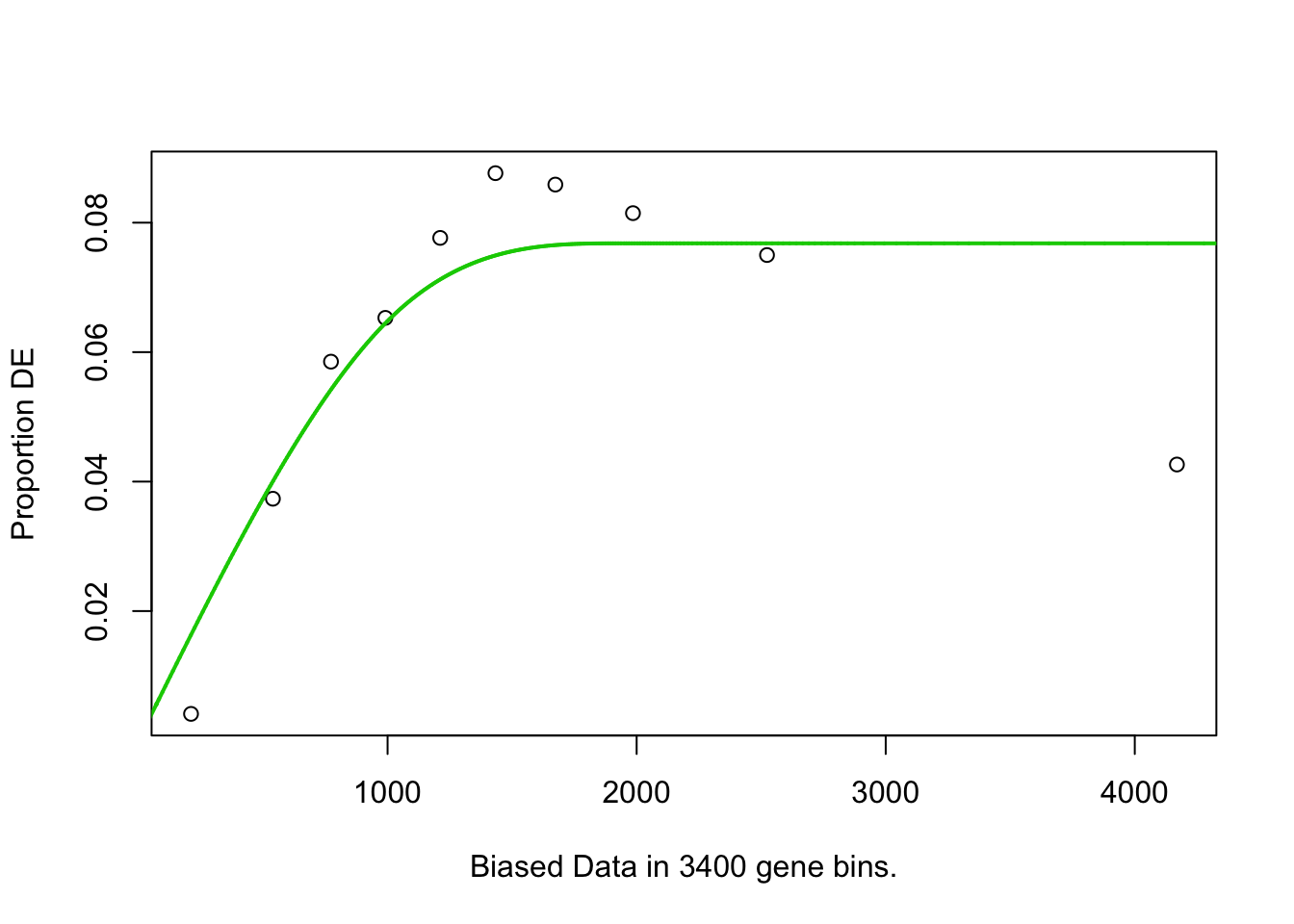
## Using manually entered categories.## Calculating the p-values...## 'select()' returned 1:1 mapping between keys and columns## [1] "enriched.GO is"
## category over_represented_pvalue under_represented_pvalue numDEInCat
## 444 GO:0006468 6.281671e-32 1.0000000 176
## 2314 GO:0042742 1.893245e-31 1.0000000 101
## 3214 GO:0071456 2.427033e-31 1.0000000 73
## 1227 GO:0010200 2.775056e-20 1.0000000 45
## 919 GO:0009617 3.128594e-18 1.0000000 40
## 997 GO:0009751 1.178078e-16 1.0000000 36
## 737 GO:0007165 8.424483e-16 1.0000000 76
## 2824 GO:0050832 9.344225e-12 1.0000000 45
## 1611 GO:0016310 2.065259e-11 1.0000000 53
## 875 GO:0009408 9.002573e-11 1.0000000 37
## 199 GO:0002237 3.175882e-09 1.0000000 15
## 2654 GO:0046777 4.913108e-09 1.0000000 39
## 3601 GO:1900426 8.907816e-09 1.0000000 13
## 201 GO:0002239 1.040633e-08 1.0000000 12
## 665 GO:0006979 1.077275e-08 1.0000000 47
## 3367 GO:0080167 5.291317e-08 1.0000000 27
## 924 GO:0009627 1.153030e-07 1.0000000 16
## 920 GO:0009620 4.738241e-07 0.9999999 22
## 3507 GO:0098542 5.059215e-07 0.9999999 14
## 2207 GO:0035556 6.907756e-07 0.9999998 34
## 915 GO:0009611 2.011069e-06 0.9999993 34
## 2938 GO:0051707 2.547530e-06 0.9999998 10
## 1863 GO:0030968 2.982841e-06 0.9999996 11
## 1185 GO:0010112 4.270211e-06 0.9999998 7
## 197 GO:0002229 4.641034e-06 0.9999989 18
## 2497 GO:0045087 7.981683e-06 0.9999983 16
## 200 GO:0002238 8.132973e-06 0.9999994 8
## 3348 GO:0080142 1.338632e-05 0.9999992 7
## 3881 GO:2000022 1.378444e-05 0.9999978 12
## 658 GO:0006952 1.404703e-05 0.9999950 28
## 738 GO:0007166 1.552398e-05 0.9999972 13
## 923 GO:0009626 1.671984e-05 0.9999970 13
## 2646 GO:0046686 1.752183e-05 0.9999922 43
## 441 GO:0006457 2.525676e-05 0.9999919 23
## 2815 GO:0050777 2.829581e-05 1.0000000 4
## 3280 GO:0072583 3.204540e-05 0.9999949 11
## 1222 GO:0010193 3.585990e-05 0.9999948 10
## 3167 GO:0071219 3.615819e-05 0.9999991 5
## 1899 GO:0031347 4.320781e-05 0.9999912 13
## 3030 GO:0060866 8.381842e-05 0.9999969 5
## 638 GO:0006886 1.125053e-04 0.9999541 29
## 203 GO:0002679 1.156134e-04 0.9999983 4
## 3889 GO:2000031 1.343777e-04 0.9999830 8
## 1476 GO:0015696 1.393526e-04 0.9999978 4
## 2706 GO:0048268 1.451030e-04 0.9999851 7
## 2952 GO:0051865 1.556018e-04 0.9999838 7
## 3921 GO:2000280 1.577103e-04 0.9999677 11
## 1051 GO:0009862 2.175134e-04 0.9999756 7
## 196 GO:0002221 2.652465e-04 0.9999768 6
## 3691 GO:1902290 2.909961e-04 0.9999824 5
## 880 GO:0009414 2.986756e-04 0.9998537 38
## 1900 GO:0031348 3.672666e-04 0.9999441 8
## 2967 GO:0052542 3.785636e-04 0.9999758 5
## 3279 GO:0072488 3.869912e-04 0.9999880 4
## 1288 GO:0010286 3.903992e-04 0.9999115 11
## 1982 GO:0032491 4.455374e-04 1.0000000 3
## 867 GO:0009311 4.500605e-04 1.0000000 3
## 29 GO:0000103 4.895936e-04 0.9999341 7
## 605 GO:0006816 6.972237e-04 0.9998631 9
## 985 GO:0009737 7.459085e-04 0.9995863 47
## 1312 GO:0010345 7.604255e-04 0.9998895 7
## numInCat
## 444 928
## 2314 391
## 3214 238
## 1227 138
## 919 118
## 997 108
## 737 411
## 2824 218
## 1611 273
## 875 169
## 199 34
## 2654 188
## 3601 27
## 201 24
## 665 301
## 3367 128
## 924 54
## 920 99
## 3507 44
## 2207 185
## 915 209
## 2938 24
## 1863 31
## 1185 13
## 197 74
## 2497 68
## 200 17
## 3348 13
## 3881 42
## 658 192
## 738 46
## 923 46
## 2646 312
## 441 138
## 2815 4
## 3280 36
## 1222 34
## 3167 7
## 1899 53
## 3030 8
## 638 204
## 203 5
## 3889 23
## 1476 5
## 2706 18
## 2952 18
## 3921 54
## 1051 19
## 196 14
## 3691 10
## 880 318
## 1900 26
## 2967 10
## 3279 6
## 1288 51
## 1982 3
## 867 3
## 29 22
## 605 34
## 985 423
## 1312 22
## term
## 444 protein phosphorylation
## 2314 defense response to bacterium
## 3214 cellular response to hypoxia
## 1227 response to chitin
## 919 response to bacterium
## 997 response to salicylic acid
## 737 signal transduction
## 2824 defense response to fungus
## 1611 phosphorylation
## 875 response to heat
## 199 response to molecule of bacterial origin
## 2654 protein autophosphorylation
## 3601 positive regulation of defense response to bacterium
## 201 response to oomycetes
## 665 response to oxidative stress
## 3367 response to karrikin
## 924 systemic acquired resistance
## 920 response to fungus
## 3507 defense response to other organism
## 2207 intracellular signal transduction
## 915 response to wounding
## 2938 response to other organism
## 1863 endoplasmic reticulum unfolded protein response
## 1185 regulation of systemic acquired resistance
## 197 defense response to oomycetes
## 2497 innate immune response
## 200 response to molecule of fungal origin
## 3348 regulation of salicylic acid biosynthetic process
## 3881 regulation of jasmonic acid mediated signaling pathway
## 658 defense response
## 738 cell surface receptor signaling pathway
## 923 plant-type hypersensitive response
## 2646 response to cadmium ion
## 441 protein folding
## 2815 negative regulation of immune response
## 3280 clathrin-dependent endocytosis
## 1222 response to ozone
## 3167 cellular response to molecule of bacterial origin
## 1899 regulation of defense response
## 3030 leaf abscission
## 638 intracellular protein transport
## 203 respiratory burst involved in defense response
## 3889 regulation of salicylic acid mediated signaling pathway
## 1476 ammonium transport
## 2706 clathrin coat assembly
## 2952 protein autoubiquitination
## 3921 regulation of root development
## 1051 systemic acquired resistance, salicylic acid mediated signaling pathway
## 196 pattern recognition receptor signaling pathway
## 3691 positive regulation of defense response to oomycetes
## 880 response to water deprivation
## 1900 negative regulation of defense response
## 2967 defense response by callose deposition
## 3279 ammonium transmembrane transport
## 1288 heat acclimation
## 1982 detection of molecule of fungal origin
## 867 oligosaccharide metabolic process
## 29 sulfate assimilation
## 605 calcium ion transport
## 985 response to abscisic acid
## 1312 suberin biosynthetic process
## ontology over_represented_padjust
## 444 BP 2.486914e-28
## 2314 BP 3.202875e-28
## 3214 BP 3.202875e-28
## 1227 BP 2.746612e-17
## 919 BP 2.477221e-15
## 997 BP 7.773353e-14
## 737 BP 4.764647e-13
## 2824 BP 4.624223e-09
## 1611 BP 9.084844e-09
## 875 BP 3.564119e-08
## 199 BP 1.143029e-06
## 2654 BP 1.620916e-06
## 3601 BP 2.712772e-06
## 201 BP 2.843287e-06
## 665 BP 2.843287e-06
## 3367 BP 1.309270e-05
## 924 BP 2.685204e-05
## 920 BP 1.042150e-04
## 3507 BP 1.054181e-04
## 2207 BP 1.367390e-04
## 915 BP 3.791344e-04
## 2938 BP 4.584396e-04
## 1863 BP 5.134377e-04
## 1185 BP 7.044069e-04
## 197 BP 7.349542e-04
## 2497 BP 1.192535e-03
## 200 BP 1.192535e-03
## 3348 BP 1.853740e-03
## 3881 BP 1.853740e-03
## 658 BP 1.853740e-03
## 738 BP 1.982563e-03
## 923 BP 2.068558e-03
## 2646 BP 2.102089e-03
## 441 BP 2.940927e-03
## 2815 BP 3.200661e-03
## 3280 BP 3.524104e-03
## 1222 BP 3.767113e-03
## 3167 BP 3.767113e-03
## 1899 BP 4.386147e-03
## 3030 BP 8.295928e-03
## 638 BP 1.086362e-02
## 203 BP 1.089794e-02
## 3889 BP 1.237212e-02
## 1476 BP 1.253857e-02
## 2706 BP 1.276584e-02
## 2952 BP 1.328458e-02
## 3921 BP 1.328458e-02
## 1051 BP 1.794032e-02
## 196 BP 2.143083e-02
## 3691 BP 2.304107e-02
## 880 BP 2.318542e-02
## 1900 BP 2.796170e-02
## 2967 BP 2.810164e-02
## 3279 BP 2.810164e-02
## 1288 BP 2.810164e-02
## 1982 BP 3.125946e-02
## 867 BP 3.125946e-02
## 29 BP 3.341899e-02
## 605 BP 4.678489e-02
## 985 BP 4.921753e-02
## 1312 BP 4.935286e-02
## named list()## Warning in pcls(G): initial point very close to some inequality constraints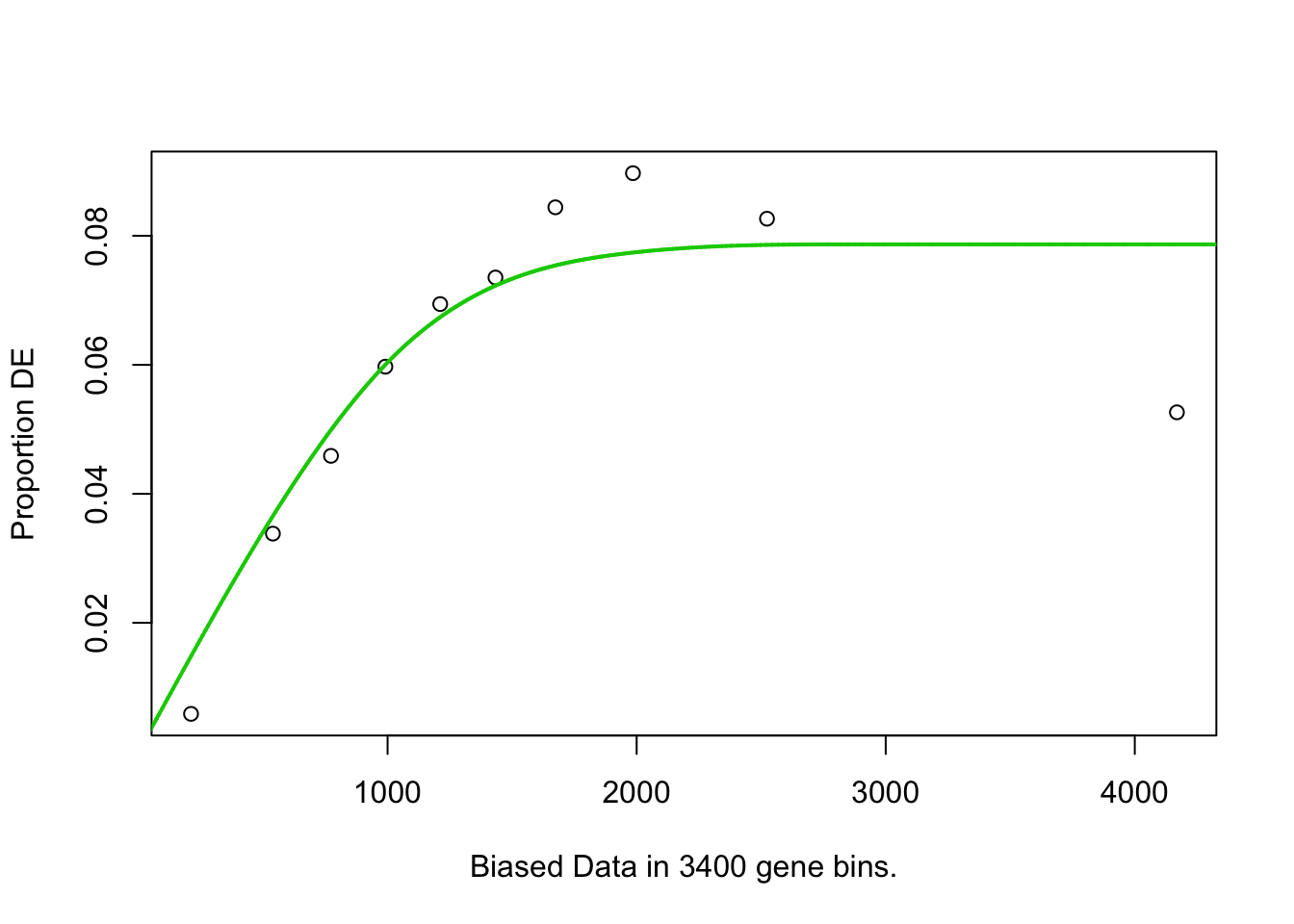
## Using manually entered categories.## Calculating the p-values...## 'select()' returned 1:1 mapping between keys and columns## [1] "enriched.GO is"
## category over_represented_pvalue under_represented_pvalue numDEInCat
## 2314 GO:0042742 6.585331e-18 1.0000000 49
## 3214 GO:0071456 1.654127e-16 1.0000000 35
## 997 GO:0009751 2.199230e-15 1.0000000 24
## 875 GO:0009408 2.355559e-13 1.0000000 27
## 919 GO:0009617 1.568629e-11 1.0000000 21
## 2869 GO:0051259 7.828841e-10 1.0000000 8
## 2299 GO:0042542 7.284227e-09 1.0000000 13
## 441 GO:0006457 3.875223e-08 1.0000000 18
## 444 GO:0006468 4.460181e-08 1.0000000 59
## 924 GO:0009627 1.371893e-07 1.0000000 11
## 2824 GO:0050832 2.677199e-07 0.9999999 22
## 737 GO:0007165 2.201645e-06 0.9999993 30
## 923 GO:0009626 7.782758e-06 0.9999991 9
## 3355 GO:0080151 8.774424e-06 0.9999999 4
## 3507 GO:0098542 2.483856e-05 0.9999972 8
## 922 GO:0009625 3.132053e-05 0.9999971 7
## 1003 GO:0009768 3.181408e-05 0.9999978 6
## 665 GO:0006979 3.353015e-05 0.9999890 22
## 1004 GO:0009769 3.446185e-05 0.9999991 4
## 658 GO:0006952 3.540787e-05 0.9999908 16
## 3889 GO:2000031 4.475935e-05 0.9999967 6
## 201 GO:0002239 4.490691e-05 0.9999967 6
## 2938 GO:0051707 5.657692e-05 0.9999956 6
## 1454 GO:0010942 8.045464e-05 0.9999976 4
## 2757 GO:0048544 1.264516e-04 0.9999851 7
## 945 GO:0009651 1.840296e-04 0.9999267 25
## 1611 GO:0016310 2.104099e-04 0.9999255 20
## numInCat term
## 2314 391 defense response to bacterium
## 3214 238 cellular response to hypoxia
## 997 108 response to salicylic acid
## 875 169 response to heat
## 919 118 response to bacterium
## 2869 17 protein complex oligomerization
## 2299 62 response to hydrogen peroxide
## 441 138 protein folding
## 444 928 protein phosphorylation
## 924 54 systemic acquired resistance
## 2824 218 defense response to fungus
## 737 411 signal transduction
## 923 46 plant-type hypersensitive response
## 3355 6 positive regulation of salicylic acid mediated signaling pathway
## 3507 44 defense response to other organism
## 922 33 response to insect
## 1003 24 photosynthesis, light harvesting in photosystem I
## 665 301 response to oxidative stress
## 1004 10 photosynthesis, light harvesting in photosystem II
## 658 192 defense response
## 3889 23 regulation of salicylic acid mediated signaling pathway
## 201 24 response to oomycetes
## 2938 24 response to other organism
## 1454 9 positive regulation of cell death
## 2757 38 recognition of pollen
## 945 395 response to salt stress
## 1611 273 phosphorylation
## ontology over_represented_padjust
## 2314 BP 2.607133e-14
## 3214 BP 3.274345e-13
## 997 BP 2.902251e-12
## 875 BP 2.331414e-10
## 919 BP 1.242041e-08
## 2869 BP 5.165730e-07
## 2299 BP 4.119751e-06
## 441 BP 1.917751e-05
## 444 BP 1.961984e-05
## 924 BP 5.431324e-05
## 2824 BP 9.635483e-05
## 737 BP 7.263593e-04
## 923 BP 2.370149e-03
## 3355 BP 2.481282e-03
## 3507 BP 6.555723e-03
## 922 BP 7.008988e-03
## 1003 BP 7.008988e-03
## 665 BP 7.008988e-03
## 1004 BP 7.008988e-03
## 658 BP 7.008988e-03
## 3889 BP 8.081202e-03
## 201 BP 8.081202e-03
## 2938 BP 9.738611e-03
## 1454 BP 1.327166e-02
## 2757 BP 2.002487e-02
## 945 BP 2.802205e-02
## 1611 BP 3.085233e-02## Warning in pcls(G): initial point very close to some inequality constraints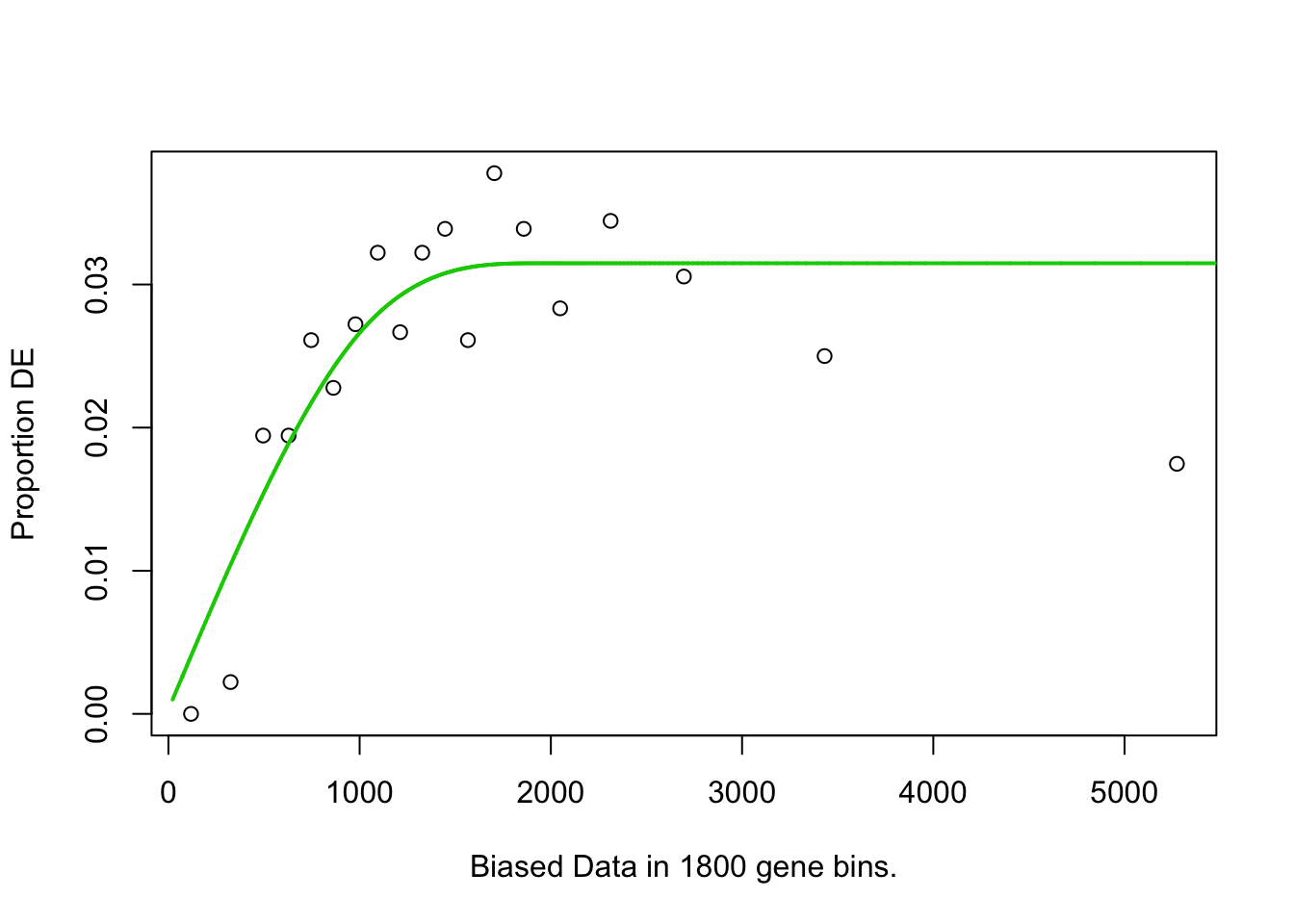
## Using manually entered categories.## Calculating the p-values...## 'select()' returned 1:1 mapping between keys and columns## Warning in pcls(G): initial point very close to some inequality constraints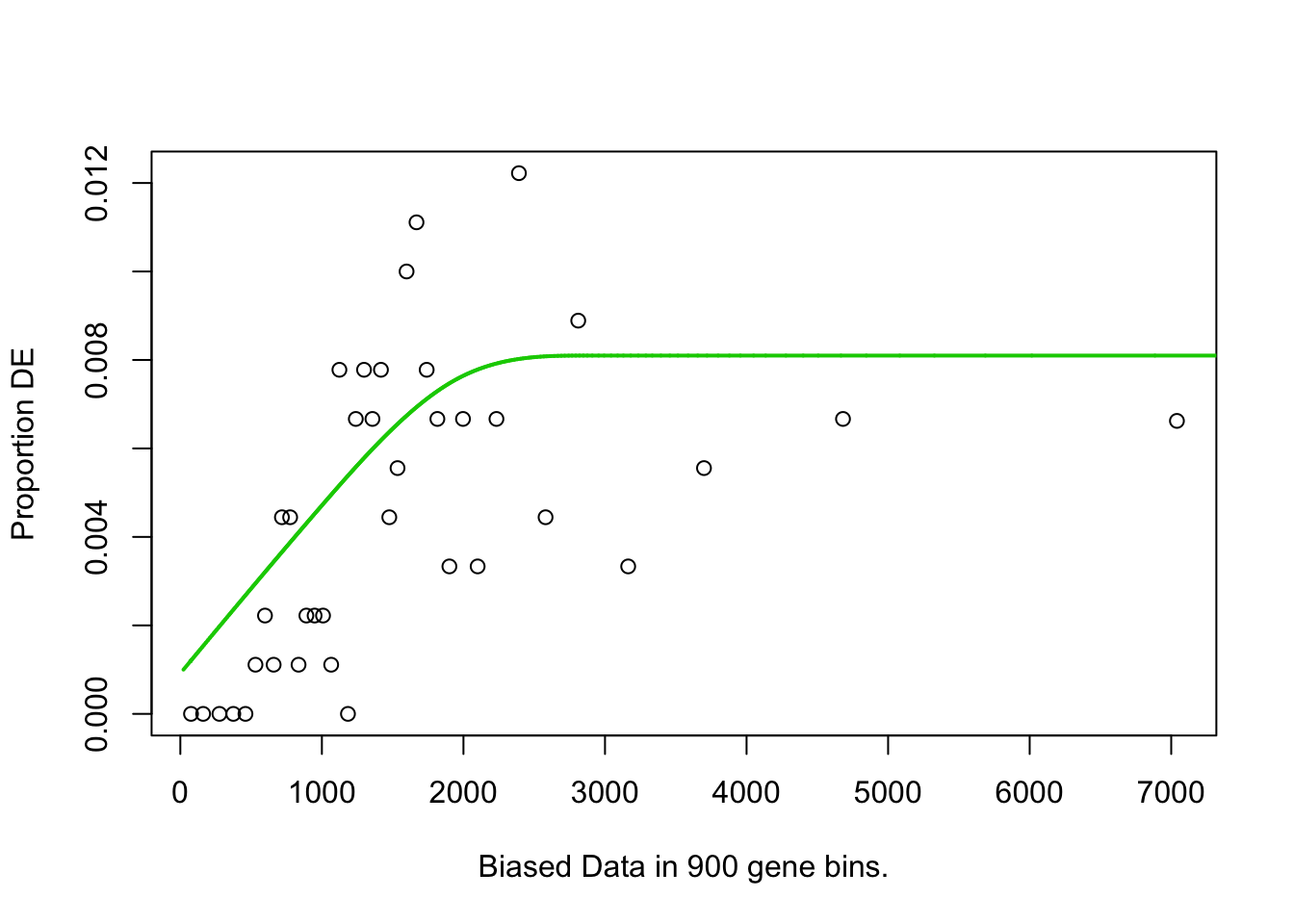
## Using manually entered categories.## Calculating the p-values...## 'select()' returned 1:1 mapping between keys and columns## [1] "enriched.GO is"
## category over_represented_pvalue under_represented_pvalue numDEInCat
## 875 GO:0009408 4.687284e-24 1.0000000 24
## 3214 GO:0071456 5.568820e-15 1.0000000 19
## 441 GO:0006457 6.355222e-14 1.0000000 15
## 2869 GO:0051259 3.022638e-12 1.0000000 7
## 2299 GO:0042542 1.572361e-11 1.0000000 10
## 2314 GO:0042742 5.448542e-08 1.0000000 16
## 945 GO:0009651 2.853867e-07 1.0000000 15
## 2938 GO:0051707 9.585648e-07 1.0000000 5
## 2141 GO:0034605 3.929501e-06 0.9999998 6
## 2144 GO:0034620 7.340712e-06 0.9999999 4
## 2840 GO:0051085 2.976695e-05 0.9999984 5
## 916 GO:0009612 4.525210e-05 0.9999994 3
## 1900 GO:0031348 4.544589e-05 0.9999985 4
## 2257 GO:0042026 1.090326e-04 0.9999953 4
## numInCat term ontology
## 875 169 response to heat BP
## 3214 238 cellular response to hypoxia BP
## 441 138 protein folding BP
## 2869 17 protein complex oligomerization BP
## 2299 62 response to hydrogen peroxide BP
## 2314 391 defense response to bacterium BP
## 945 395 response to salt stress BP
## 2938 24 response to other organism BP
## 2141 59 cellular response to heat BP
## 2144 17 cellular response to unfolded protein BP
## 2840 48 chaperone cofactor-dependent protein refolding BP
## 916 10 response to mechanical stimulus BP
## 1900 26 negative regulation of defense response BP
## 2257 32 protein refolding BP
## over_represented_padjust
## 875 1.855696e-20
## 3214 1.102348e-11
## 441 8.386775e-11
## 2869 2.991656e-09
## 2299 1.244995e-08
## 2314 3.595130e-05
## 945 1.614065e-04
## 2938 4.743698e-04
## 2141 1.728544e-03
## 2144 2.906188e-03
## 2840 1.071340e-02
## 916 1.384002e-02
## 1900 1.384002e-02
## 2257 3.083287e-02## Warning in pcls(G): initial point very close to some inequality constraints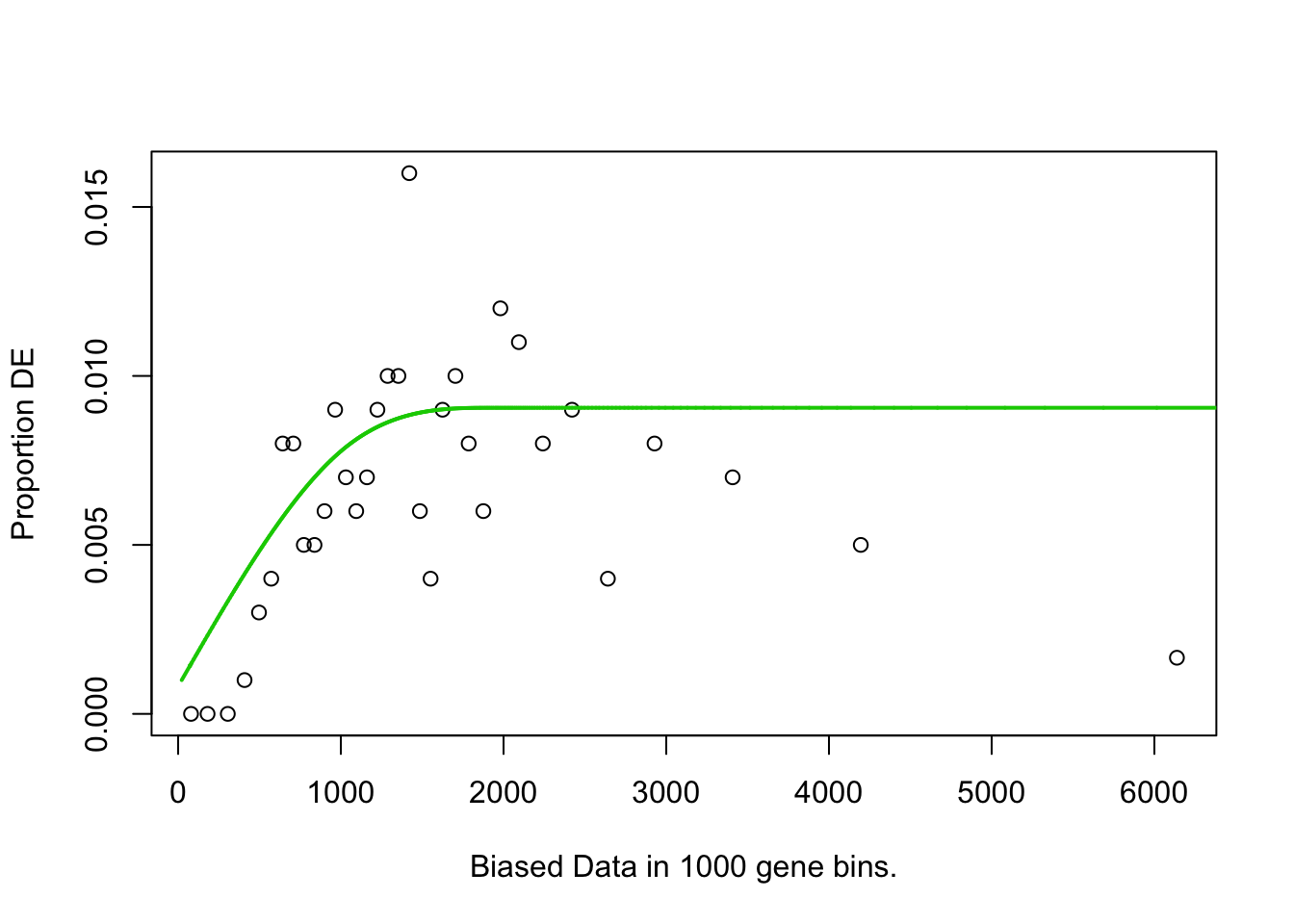
## Using manually entered categories.## Calculating the p-values...## 'select()' returned 1:1 mapping between keys and columns## [1] "enriched.GO is"
## category over_represented_pvalue under_represented_pvalue numDEInCat
## 1003 GO:0009768 3.548079e-06 0.9999999 4
## 1004 GO:0009769 5.987083e-06 1.0000000 3
## numInCat term ontology
## 1003 24 photosynthesis, light harvesting in photosystem I BP
## 1004 10 photosynthesis, light harvesting in photosystem II BP
## over_represented_padjust
## 1003 0.01185143
## 1004 0.01185143## Warning in pcls(G): initial point very close to some inequality constraints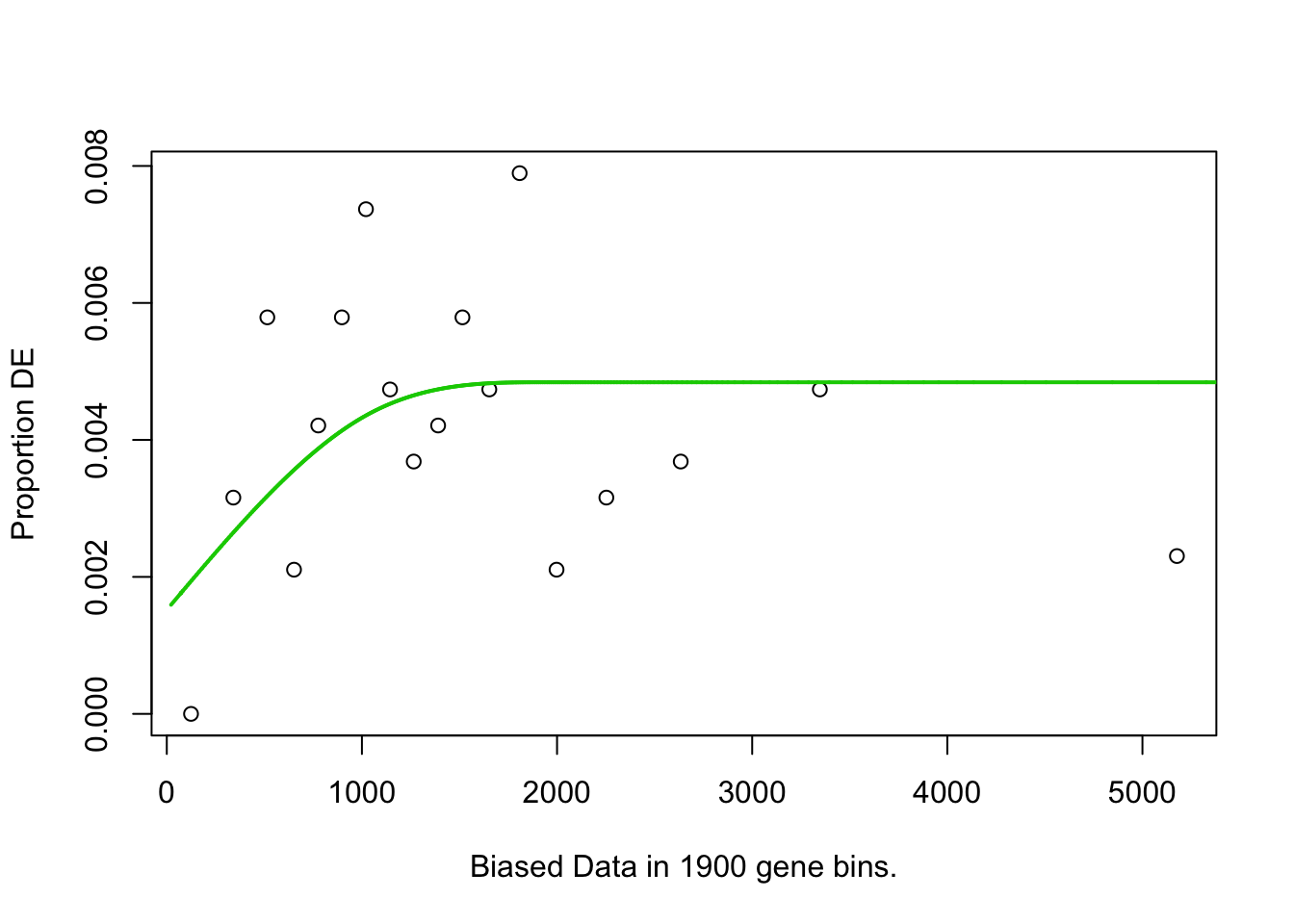
## Using manually entered categories.## Calculating the p-values...## 'select()' returned 1:1 mapping between keys and columns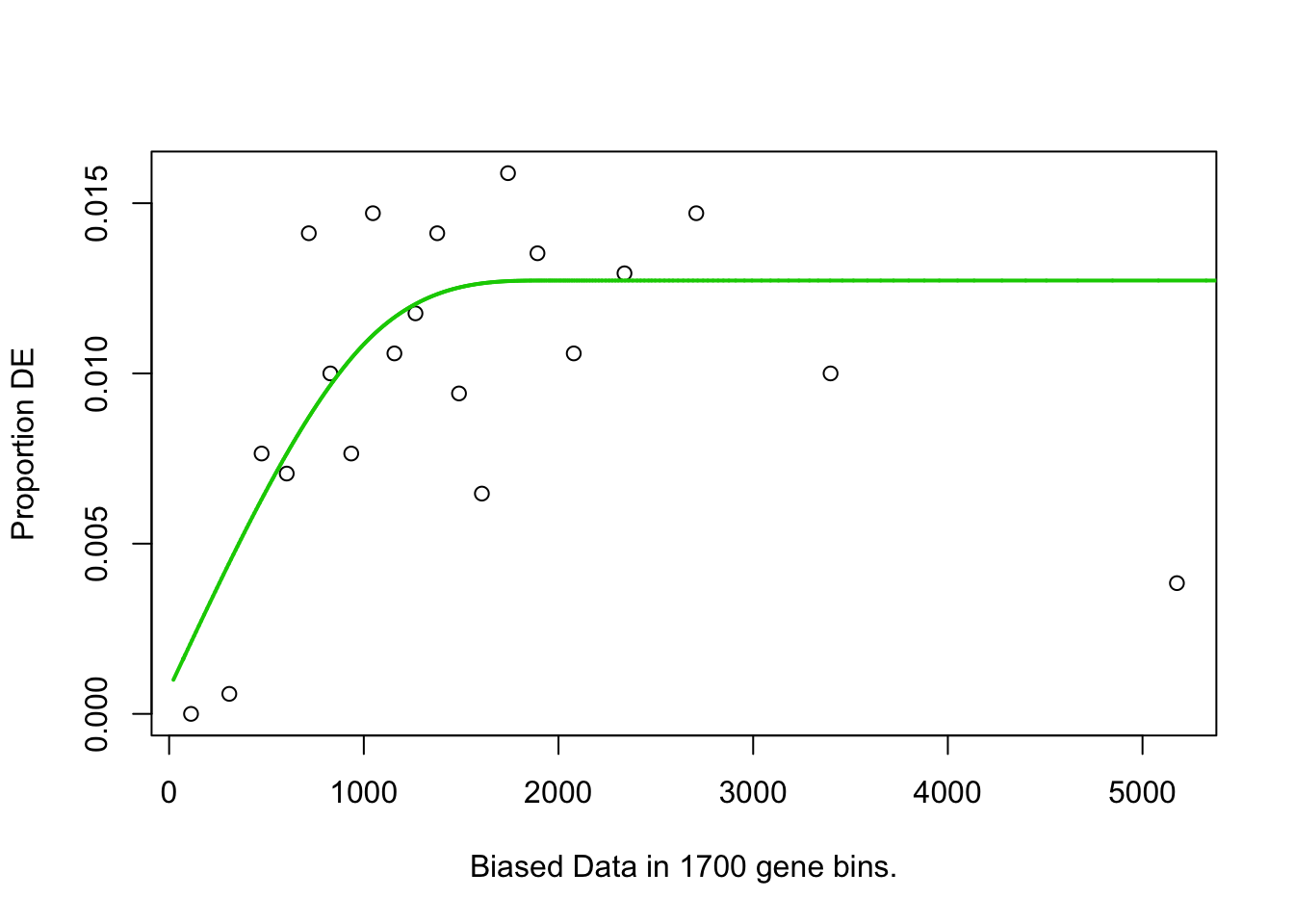
## [1] "enriched.GO is"
## category over_represented_pvalue under_represented_pvalue numDEInCat
## 997 GO:0009751 4.961497e-14 1.0000000 16
## 2314 GO:0042742 3.274247e-12 1.0000000 25
## 444 GO:0006468 7.885881e-11 1.0000000 37
## 737 GO:0007165 1.638219e-07 1.0000000 19
## 919 GO:0009617 7.931019e-07 0.9999999 10
## 3214 GO:0071456 1.336782e-06 0.9999998 13
## 2654 GO:0046777 2.745446e-06 0.9999996 12
## 1611 GO:0016310 3.975974e-06 0.9999992 14
## 201 GO:0002239 5.727640e-06 0.9999998 5
## 197 GO:0002229 2.517999e-05 0.9999976 7
## 2824 GO:0050832 3.146979e-05 0.9999943 11
## 3507 GO:0098542 1.097841e-04 0.9999924 5
## numInCat term ontology
## 997 108 response to salicylic acid BP
## 2314 391 defense response to bacterium BP
## 444 928 protein phosphorylation BP
## 737 411 signal transduction BP
## 919 118 response to bacterium BP
## 3214 238 cellular response to hypoxia BP
## 2654 188 protein autophosphorylation BP
## 1611 273 phosphorylation BP
## 201 24 response to oomycetes BP
## 197 74 defense response to oomycetes BP
## 2824 218 defense response to fungus BP
## 3507 44 defense response to other organism BP
## over_represented_padjust
## 997 1.964257e-10
## 2314 6.481371e-09
## 444 1.040673e-07
## 737 1.621427e-04
## 919 6.279780e-04
## 3214 8.820533e-04
## 2654 1.552746e-03
## 1611 1.967610e-03
## 201 2.519525e-03
## 197 9.968757e-03
## 2824 1.132626e-02
## 3507 3.621962e-02
## $GO.ORA.temp.up_up
## [1] "no enriched GO"read GOseq result table
eGOseqs<-list.files(pattern="enrich.Rdata",path=file.path("output"))
eGOseqs.list2<-sapply(file.path("output",eGOseqs),function(x) mget(load(x))) # mget will return the value of the object(or objects) in a list. see https://stackoverflow.com/questions/29398630/load-data-frames-into-list
#names(eGOseqs.list2)
eGOseqs.list2.summary<-do.call("rbind",eGOseqs.list2)
#head(eGOseqs.list2.summary) # make sure those are file names
rownames(eGOseqs.list2.summary)<-1:nrow(eGOseqs.list2.summary)
View(eGOseqs.list2.summary)Drawing heatmap
library(scales) # for mute in plot##
## Attaching package: 'scales'## The following object is masked from 'package:purrr':
##
## discard## The following object is masked from 'package:readr':
##
## col_factor# eGOseqs.list2.summary (more than three genes in each KO)
eGOseqs.list2.summary<-eGOseqs.list2.summary[eGOseqs.list2.summary$numDEInCat>3,]
# focused on only very significant GO terms
eGOseqs.list2.summary<-eGOseqs.list2.summary[eGOseqs.list2.summary$over_represented_padjust<1e-4,]
# how to cluster GO Ontology according to this pattern?
## using hclust to sort GO terms
GO.list<-unique(eGOseqs.list2.summary$category)
sample.list<-unique(eGOseqs.list2.summary$sample_FC) # revised
# having x-label also used the x-label for df below
eGOseqs.list2.summary <- eGOseqs.list2.summary %>% mutate(sample=str_remove(sample,".csv")) %>% mutate(sample=str_remove(sample,".DEGs.int")) %>% unite(sample_FC,sample,FC)
# Making matrix for calculate hierarchical clustering of "over-representd_padjust" value using sperad
df<-eGOseqs.list2.summary %>% dplyr::select(category,sample_FC,over_represented_padjust) %>% spread(sample_FC,over_represented_padjust,-1)
df[df<1e-100]<-1e-100 # to avoid "Inf" after log10 transformation
df<- df %>% mutate_at(vars(2:11) ,.funs=function(x) -log10(x))
# df<- df %>% mutate_at(vars(2:11) ,.funs=-log10) # does not work
df[is.na(df)]<-1
df2<-df[,2:11]
rownames(df2)<-df$category
hc<-stats::hclust(dist(df2), "ave") # only numbers
hc$order## [1] 23 66 68 7 58 73 43 46 61 47 42 60 17 41 36 33 48 24 6 21 32 26 40 49 35
## [26] 4 51 29 38 14 30 25 27 10 11 50 12 13 18 22 9 55 62 54 67 3 57 56 20 64
## [51] 71 34 1 52 37 44 70 53 5 15 59 8 39 31 28 65 45 16 72 19 69 2 63hc.tib<-tibble(category=hc$labels,order=hc$order)
# change term order using hclust() results
hc.tib2<-hc.tib %>% inner_join(eGOseqs.list2.summary[,c("category","term")],by="category") %>% distinct() %>% arrange(desc(order))
eGOseqs.list2.summary$term<-factor(eGOseqs.list2.summary$term,levels=as_vector(hc.tib2$term))
# To avoid gray color, add 1e-20 to smaller padjust value
eGOseqs.list2.summary$over_represented_padjust[eGOseqs.list2.summary$over_represented_padjust<1e-50]<-1e-50
# format x label
#eGOseqs.list2.summary <- eGOseqs.list2.summary %>% mutate(sample2=str_replace(sample,"trt.density","trt_density")) %>% separate(sample2,into=c("tissue","factor","DEGs","model","ref_density","ref_soil"),sep="\\.") %>% mutate(ref_soil=str_remove(ref_soil,"r")) %>% mutate(ref_density=str_remove(ref_density,"r")) %>% unite(density_FC,ref_density,FC) # simplify elements in "density_FC" column for simpler x label in heatmap
# plot
GOseq.plot<-ggplot(eGOseqs.list2.summary,aes(x=sample_FC,y=term)) + geom_tile(aes(fill=-log10(over_represented_padjust)),colour="black") + scale_fill_gradient2(limit=c(0,50),low=muted("green"), high=muted("magenta")) # OK
GOseq.plot<-GOseq.plot+ theme(axis.text.x=element_text(size=10,angle=90),
axis.text.y=element_text(size=10),
axis.title=element_text(size=10),
axis.ticks = element_blank(),
panel.background = element_rect(fill = "white",colour="black"),
plot.title=element_text(size=20),
axis.line=element_blank()) + labs(x="",y="",title="",fill="-log10\n pvalue")
GOseq.plot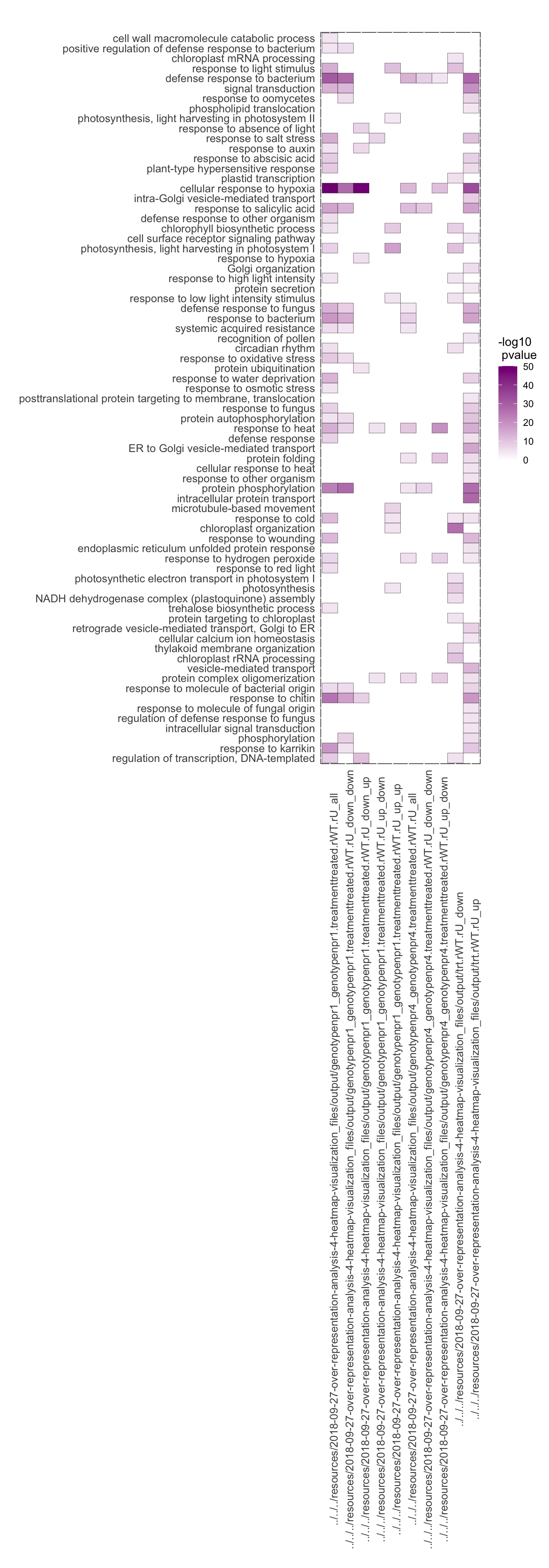
# ggplot_build(GOseq.plot)Session info
sessionInfo()## R version 3.6.2 (2019-12-12)
## Platform: x86_64-apple-darwin15.6.0 (64-bit)
## Running under: macOS Mojave 10.14.6
##
## Matrix products: default
## BLAS: /Library/Frameworks/R.framework/Versions/3.6/Resources/lib/libRblas.0.dylib
## LAPACK: /Library/Frameworks/R.framework/Versions/3.6/Resources/lib/libRlapack.dylib
##
## locale:
## [1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
##
## attached base packages:
## [1] stats4 parallel stats graphics grDevices utils datasets
## [8] methods base
##
## other attached packages:
## [1] scales_1.1.1 annotate_1.64.0
## [3] XML_3.99-0.3 GO.db_3.10.0
## [5] AnnotationDbi_1.48.0 goseq_1.38.0
## [7] geneLenDataBase_1.22.0 BiasedUrn_1.07
## [9] ShortRead_1.44.3 GenomicAlignments_1.22.1
## [11] SummarizedExperiment_1.16.1 DelayedArray_0.12.3
## [13] matrixStats_0.57.0 Biobase_2.46.0
## [15] Rsamtools_2.2.3 GenomicRanges_1.38.0
## [17] GenomeInfoDb_1.22.1 Biostrings_2.54.0
## [19] XVector_0.26.0 IRanges_2.20.2
## [21] S4Vectors_0.24.4 BiocParallel_1.20.1
## [23] BiocGenerics_0.32.0 forcats_0.5.0
## [25] stringr_1.4.0 dplyr_1.0.2
## [27] purrr_0.3.4 readr_1.4.0
## [29] tidyr_1.1.2 tibble_3.0.4
## [31] ggplot2_3.3.3 tidyverse_1.3.0
##
## loaded via a namespace (and not attached):
## [1] colorspace_2.0-0 hwriter_1.3.2 ellipsis_0.3.1
## [4] fs_1.5.0 rstudioapi_0.13 farver_2.0.3
## [7] bit64_4.0.5 fansi_0.4.1 lubridate_1.7.9.2
## [10] xml2_1.3.2 splines_3.6.2 knitr_1.31
## [13] jsonlite_1.7.2 broom_0.7.3 dbplyr_2.0.0
## [16] png_0.1-7 compiler_3.6.2 httr_1.4.2
## [19] backports_1.2.1 assertthat_0.2.1 Matrix_1.3-2
## [22] cli_2.2.0 htmltools_0.5.1.1 prettyunits_1.1.1
## [25] tools_3.6.2 gtable_0.3.0 glue_1.4.2
## [28] GenomeInfoDbData_1.2.2 rappdirs_0.3.1 Rcpp_1.0.6
## [31] cellranger_1.1.0 vctrs_0.3.6 nlme_3.1-151
## [34] blogdown_1.2.2 rtracklayer_1.46.0 xfun_0.22
## [37] rvest_0.3.6 lifecycle_0.2.0 zlibbioc_1.32.0
## [40] hms_0.5.3 RColorBrewer_1.1-2 yaml_2.2.1
## [43] curl_4.3 memoise_1.1.0 biomaRt_2.42.1
## [46] latticeExtra_0.6-29 stringi_1.5.3 RSQLite_2.2.1
## [49] highr_0.8 GenomicFeatures_1.38.2 rlang_0.4.10
## [52] pkgconfig_2.0.3 bitops_1.0-6 evaluate_0.14
## [55] lattice_0.20-41 labeling_0.4.2 bit_4.0.4
## [58] tidyselect_1.1.0 magrittr_2.0.1 bookdown_0.21
## [61] R6_2.5.0 generics_0.1.0 DBI_1.1.0
## [64] pillar_1.4.7 haven_2.3.1 withr_2.3.0
## [67] mgcv_1.8-33 RCurl_1.98-1.2 modelr_0.1.8
## [70] crayon_1.3.4 utf8_1.1.4 BiocFileCache_1.10.2
## [73] rmarkdown_2.7 jpeg_0.1-8.1 progress_1.2.2
## [76] grid_3.6.2 readxl_1.3.1 blob_1.2.1
## [79] reprex_0.3.0 digest_0.6.27 xtable_1.8-4
## [82] openssl_1.4.3 munsell_0.5.0 askpass_1.1